stat_debug_group and stat_debug_panel
apply a function to data and, most importantly,
echo to the R console their data input or a summary of it.
Usage
stat_debug_group(
mapping = NULL,
data = NULL,
geom = "null",
fun.data = "I",
fun.data.args = list(),
dbgfun.data = "head",
dbgfun.data.args = list(),
geom.dbgfun.data = "head",
geom.dbgfun.data.args = list(),
geom.dbgfun.params = NULL,
geom.dbgfun.params.args = list(),
dbgfun.print = "print",
dbgfun.print.args = list(),
position = "identity",
na.rm = FALSE,
show.legend = FALSE,
inherit.aes = TRUE,
...
)
stat_debug_panel(
mapping = NULL,
data = NULL,
geom = "null",
fun.data = "I",
fun.data.args = list(),
dbgfun.data = "head",
dbgfun.data.args = list(),
geom.dbgfun.data = "head",
geom.dbgfun.data.args = list(),
geom.dbgfun.params = NULL,
geom.dbgfun.params.args = list(),
dbgfun.print = "print",
dbgfun.print.args = list(),
position = "identity",
na.rm = FALSE,
show.legend = FALSE,
inherit.aes = TRUE,
...
)Arguments
- mapping
The aesthetic mapping, usually constructed with
aesoraes_. Only needs to be set at the layer level if you are overriding the plot defaults.- data
A layer specific dataset - only needed if you want to override the plot defaults.
- geom
The geometric object to use display the data
- fun.data
A function taking a data frame as its first argument and returning a data frame. This function does the computations generating the value passed from the statistic to the downstream geometry.
- fun.data.args
A named list of additional arguments to be passed to
fun.data.- dbgfun.data, geom.dbgfun.data, geom.dbgfun.params
A functions used to summarise the
dataandparametersobjects received as input by the statistic and geometry.- dbgfun.data.args, geom.dbgfun.data.args, geom.dbgfun.params.args
A named list of arguments.
- dbgfun.print
A function used to print the summary of the
dataobject received as input by the statistic, also visible to the geometry, and used bygeom_debug().- dbgfun.print.args
A named list. Currently ignored!
- position
The position adjustment to use for overlapping points on this layer
- na.rm
a logical value indicating whether NA values should be stripped before the computation proceeds.
- show.legend
logical. Should this layer be included in the legends?
NA, the default, includes if any aesthetics are mapped.FALSEnever includes, andTRUEalways includes.- inherit.aes
If
FALSE, overrides the default aesthetics, rather than combining with them. This is most useful for helper functions that define both data and aesthetics and shouldn't inherit behaviour from the default plot specification, e.g.borders.- ...
other arguments passed on to
layer. This can include aesthetics whose values you want to set, not map. Seelayerfor more details.
Value
A copy of its data input, which is an object of class
"data.frame" or inheriting from "data.frame".
Details
These stats are meant to be used for the side-effect of printing to
the console the data object received as input by the
compute_group() or compute_panel() function, or a summary of
it. These data objects are the same as those received as input by
any other statistics passed the same arguments. By default, the applied
function is I(), the identity function.
In principle any geom can be passed as argument to override geom =
"null". However, geom = "debug_panel" and geom =
"debug_group" are treated as special cases and functions
geom.dbgfun.data and geom.dbgfun.params, and lists
geom.dbgfun.data.args and geom.dbgfun.params.args renamed and
passed to the geometry. Arguments passed to these four formal parameters
are not passed to other geometries.
Keep in mind that this stat sets default mappings only for the x
and/or y aesthetics, additional mappings can be set using
aes(), possibly together with after_stat().
Computed variables
- x
x at centre of range
- y
y at centre of range
- nrow
nrow()ofdataobject- ncol
ncol()ofdataobject- colnames
colnames()ofdataobject- colclasses
class()ofxandycolumns indataobject- group
all distinct values in group as passed in
dataobject- PANEL
all distinct values in PANEL as passed in
dataobject
Examples
my.df <- data.frame(x = rep(1:10, 2),
y = rep(c(1,2), c(10,10)) + rnorm(20),
group = rep(c("A","B"), c(10,10)))
# by default head() is used to show the top rows of the data object
# and geom_null() to silence the data returned by the stat
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group()
 #> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
# geom_debug prints the data returned by the stat
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(geom = "debug_panel")
#> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
# geom_debug prints the data returned by the stat
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(geom = "debug_panel")
 #> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
# geom_debug prints the data returned by the stat
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(geom = "debug_panel",
geom.dbgfun.params = "summary")
#> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
# geom_debug prints the data returned by the stat
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(geom = "debug_panel",
geom.dbgfun.params = "summary")
 #> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'params' (summary):"
#> Length Class Mode
#> x 11 ViewScale environment
#> x.sec 11 ViewScale environment
#> x.range 2 -none- numeric
#> y 11 ViewScale environment
#> y.sec 11 ViewScale environment
#> y.range 2 -none- numeric
#> reverse 1 -none- character
#> guides 4 Guides environment
# to print only the the data returned by the stat
# we pass as summary function a function that always returns NULL
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(geom = "debug_panel",
dbgfun.data = function(x) {NULL})
#> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'params' (summary):"
#> Length Class Mode
#> x 11 ViewScale environment
#> x.sec 11 ViewScale environment
#> x.range 2 -none- numeric
#> y 11 ViewScale environment
#> y.sec 11 ViewScale environment
#> y.range 2 -none- numeric
#> reverse 1 -none- character
#> guides 4 Guides environment
# to print only the the data returned by the stat
# we pass as summary function a function that always returns NULL
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(geom = "debug_panel",
dbgfun.data = function(x) {NULL})
 #> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (anonymous function):"
#> NULL
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(aes(label = paste("group:", group)),
geom = "text")
#> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (anonymous function):"
#> NULL
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(aes(label = paste("group:", group)),
geom = "text")
 #> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> label x y PANEL group
#> 1 group: A 1 -0.9100875 1 -1
#> 2 group: A 2 0.7207628 1 -1
#> 3 group: A 3 0.6865540 1 -1
#> 4 group: A 4 2.0673079 1 -1
#> 5 group: A 5 1.0700349 1 -1
#> 6 group: A 6 0.3608767 1 -1
# here we show all the data object
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(dbgfun.data = "I")
#> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> label x y PANEL group
#> 1 group: A 1 -0.9100875 1 -1
#> 2 group: A 2 0.7207628 1 -1
#> 3 group: A 3 0.6865540 1 -1
#> 4 group: A 4 2.0673079 1 -1
#> 5 group: A 5 1.0700349 1 -1
#> 6 group: A 6 0.3608767 1 -1
# here we show all the data object
ggplot(my.df, aes(x,y)) +
geom_point() +
stat_debug_group(dbgfun.data = "I")
 #> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (I):"
#> x y PANEL group
#> 1 1 -0.91008747 1 -1
#> 2 2 0.72076276 1 -1
#> 3 3 0.68655402 1 -1
#> 4 4 2.06730788 1 -1
#> 5 5 1.07003485 1 -1
#> 6 6 0.36087668 1 -1
#> 7 7 0.95003510 1 -1
#> 8 8 0.74851656 1 -1
#> 9 9 1.44479712 1 -1
#> 10 10 3.75541758 1 -1
#> 11 1 2.04653138 1 -1
#> 12 2 2.57770907 1 -1
#> 13 3 2.11819487 1 -1
#> 14 4 0.08827951 1 -1
#> 15 5 2.86208648 1 -1
#> 16 6 1.75676326 1 -1
#> 17 7 1.79391281 1 -1
#> 18 8 2.01917759 1 -1
#> 19 9 2.02956075 1 -1
#> 20 10 2.54982754 1 -1
# with grouping
ggplot(my.df, aes(x,y, colour = group)) +
geom_point() +
stat_debug_group()
#> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (I):"
#> x y PANEL group
#> 1 1 -0.91008747 1 -1
#> 2 2 0.72076276 1 -1
#> 3 3 0.68655402 1 -1
#> 4 4 2.06730788 1 -1
#> 5 5 1.07003485 1 -1
#> 6 6 0.36087668 1 -1
#> 7 7 0.95003510 1 -1
#> 8 8 0.74851656 1 -1
#> 9 9 1.44479712 1 -1
#> 10 10 3.75541758 1 -1
#> 11 1 2.04653138 1 -1
#> 12 2 2.57770907 1 -1
#> 13 3 2.11819487 1 -1
#> 14 4 0.08827951 1 -1
#> 15 5 2.86208648 1 -1
#> 16 6 1.75676326 1 -1
#> 17 7 1.79391281 1 -1
#> 18 8 2.01917759 1 -1
#> 19 9 2.02956075 1 -1
#> 20 10 2.54982754 1 -1
# with grouping
ggplot(my.df, aes(x,y, colour = group)) +
geom_point() +
stat_debug_group()
 #> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (head):"
#> x y colour PANEL group
#> 1 1 -0.9100875 A 1 1
#> 2 2 0.7207628 A 1 1
#> 3 3 0.6865540 A 1 1
#> 4 4 2.0673079 A 1 1
#> 5 5 1.0700349 A 1 1
#> 6 6 0.3608767 A 1 1
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (head):"
#> x y colour PANEL group
#> 11 1 2.04653138 B 1 2
#> 12 2 2.57770907 B 1 2
#> 13 3 2.11819487 B 1 2
#> 14 4 0.08827951 B 1 2
#> 15 5 2.86208648 B 1 2
#> 16 6 1.75676326 B 1 2
ggplot(my.df, aes(x,y, colour = group)) +
geom_point() +
stat_debug_panel()
#> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (head):"
#> x y colour PANEL group
#> 1 1 -0.9100875 A 1 1
#> 2 2 0.7207628 A 1 1
#> 3 3 0.6865540 A 1 1
#> 4 4 2.0673079 A 1 1
#> 5 5 1.0700349 A 1 1
#> 6 6 0.3608767 A 1 1
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (head):"
#> x y colour PANEL group
#> 11 1 2.04653138 B 1 2
#> 12 2 2.57770907 B 1 2
#> 13 3 2.11819487 B 1 2
#> 14 4 0.08827951 B 1 2
#> 15 5 2.86208648 B 1 2
#> 16 6 1.75676326 B 1 2
ggplot(my.df, aes(x,y, colour = group)) +
geom_point() +
stat_debug_panel()
 #> [1] "PANEL 1; group(s) 1, 2; 'compute_panel()' input 'data' (head):"
#> x y colour PANEL group
#> 1 1 -0.9100875 A 1 1
#> 2 2 0.7207628 A 1 1
#> 3 3 0.6865540 A 1 1
#> 4 4 2.0673079 A 1 1
#> 5 5 1.0700349 A 1 1
#> 6 6 0.3608767 A 1 1
ggplot(my.df, aes(x, y, colour = group)) +
geom_point() +
stat_debug_group(dbgfun.data = "nrow")
#> [1] "PANEL 1; group(s) 1, 2; 'compute_panel()' input 'data' (head):"
#> x y colour PANEL group
#> 1 1 -0.9100875 A 1 1
#> 2 2 0.7207628 A 1 1
#> 3 3 0.6865540 A 1 1
#> 4 4 2.0673079 A 1 1
#> 5 5 1.0700349 A 1 1
#> 6 6 0.3608767 A 1 1
ggplot(my.df, aes(x, y, colour = group)) +
geom_point() +
stat_debug_group(dbgfun.data = "nrow")
 #> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (nrow):"
#> [1] 10
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (nrow):"
#> [1] 10
ggplot(my.df, aes(x, y)) +
geom_point() +
facet_wrap(~group) +
stat_debug_group()
#> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (nrow):"
#> [1] 10
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (nrow):"
#> [1] 10
ggplot(my.df, aes(x, y)) +
geom_point() +
facet_wrap(~group) +
stat_debug_group()
 #> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 2; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 2.04653138 2 -1
#> 2 2 2.57770907 2 -1
#> 3 3 2.11819487 2 -1
#> 4 4 0.08827951 2 -1
#> 5 5 2.86208648 2 -1
#> 6 6 1.75676326 2 -1
# by default head() is used to show the top rows of data object
ggplot(my.df, aes(group,y)) +
geom_point() +
stat_debug_group()
#> [1] "PANEL 1; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 -1
#> 2 2 0.7207628 1 -1
#> 3 3 0.6865540 1 -1
#> 4 4 2.0673079 1 -1
#> 5 5 1.0700349 1 -1
#> 6 6 0.3608767 1 -1
#> [1] "PANEL 2; group(s) -1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 2.04653138 2 -1
#> 2 2 2.57770907 2 -1
#> 3 3 2.11819487 2 -1
#> 4 4 0.08827951 2 -1
#> 5 5 2.86208648 2 -1
#> 6 6 1.75676326 2 -1
# by default head() is used to show the top rows of data object
ggplot(my.df, aes(group,y)) +
geom_point() +
stat_debug_group()
 #> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 1
#> 2 1 0.7207628 1 1
#> 3 1 0.6865540 1 1
#> 4 1 2.0673079 1 1
#> 5 1 1.0700349 1 1
#> 6 1 0.3608767 1 1
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 11 2 2.04653138 1 2
#> 12 2 2.57770907 1 2
#> 13 2 2.11819487 1 2
#> 14 2 0.08827951 1 2
#> 15 2 2.86208648 1 2
#> 16 2 1.75676326 1 2
ggplot(my.df, aes(group,y)) +
geom_point() +
stat_debug_group(dbgfun.data.args = list(n = 3))
#> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 1
#> 2 1 0.7207628 1 1
#> 3 1 0.6865540 1 1
#> 4 1 2.0673079 1 1
#> 5 1 1.0700349 1 1
#> 6 1 0.3608767 1 1
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 11 2 2.04653138 1 2
#> 12 2 2.57770907 1 2
#> 13 2 2.11819487 1 2
#> 14 2 0.08827951 1 2
#> 15 2 2.86208648 1 2
#> 16 2 1.75676326 1 2
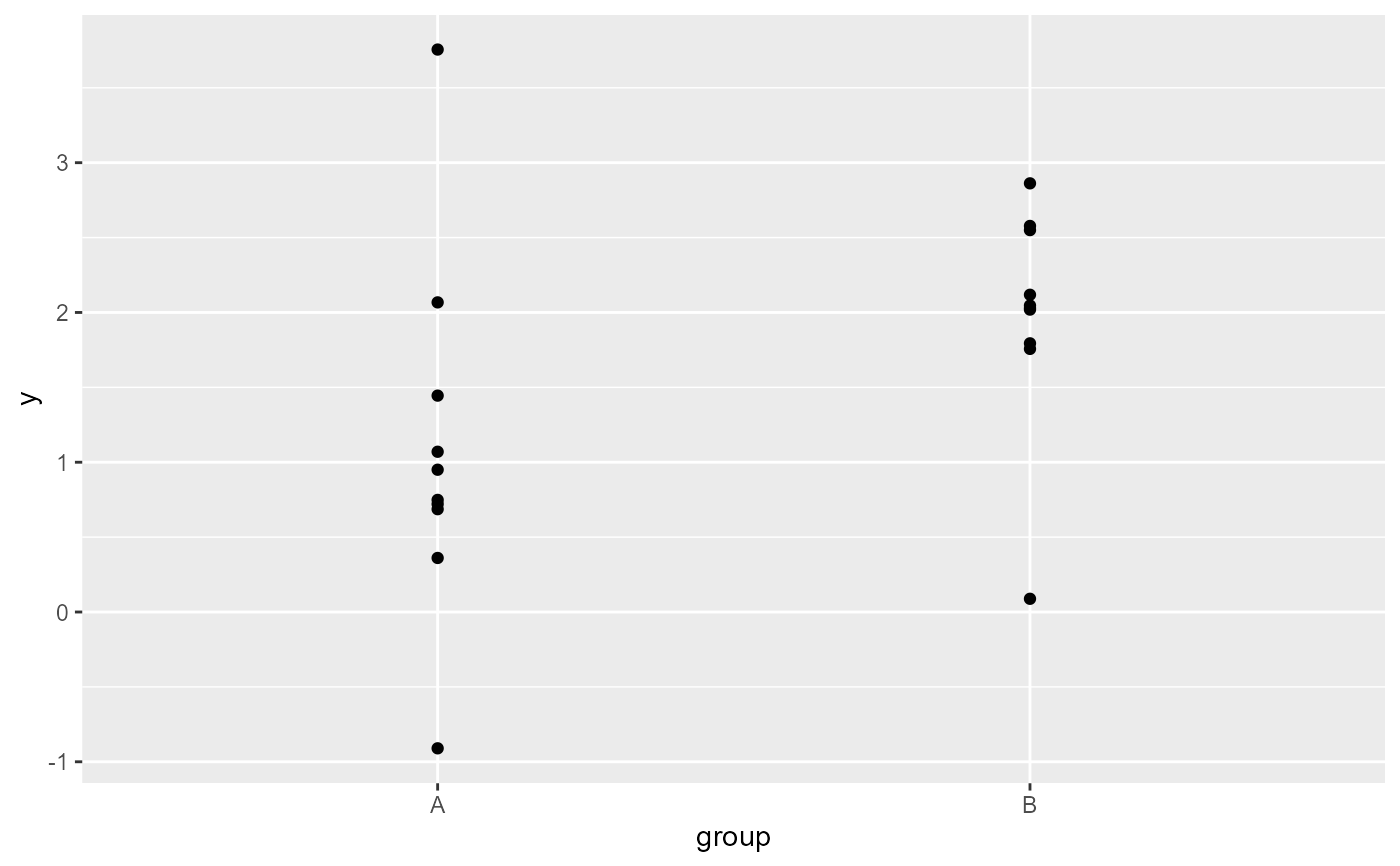
ggplot(my.df, aes(group,y)) +
geom_point() +
stat_debug_group(dbgfun.data.args = list(n = 3))
 #> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 1
#> 2 1 0.7207628 1 1
#> 3 1 0.6865540 1 1
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 11 2 2.046531 1 2
#> 12 2 2.577709 1 2
#> 13 2 2.118195 1 2
#> [1] "PANEL 1; group(s) 1; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 1 1 -0.9100875 1 1
#> 2 1 0.7207628 1 1
#> 3 1 0.6865540 1 1
#> [1] "PANEL 1; group(s) 2; 'compute_group()' input 'data' (head):"
#> x y PANEL group
#> 11 2 2.046531 1 2
#> 12 2 2.577709 1 2
#> 13 2 2.118195 1 2
