
User Guide: 1 Using the Data
Package ‘photobiologySensors’ 0.5.3
Pedro J. Aphalo
2026-03-13
Source:vignettes/userguide-1.Rmd
userguide-1.RmdIntroduction
This package, is a data only package, part of a suite, which has package ‘photobiology’ at its core. Please visit (https://www.r4photobiology.info/) for more details. For more details on plotting spectra, please consult the documentation for package ‘ggspectra’, and for information on the calculation of summaries and maths operations between spectra, please, consult the documentation for package ‘photobiology’.
library(photobiology)
library(photobiologySensors)
library(photobiologyWavebands)
eval_ggspectra <- TRUE
library(ggspectra)In this brief User Guide we describe how to re-scale the normalized spectra, and how to access individual spectra or subsets of spectra.
Data and metadata
Spectra are contained in two collection: sensors.mspct
contains spectral data for various types of broadband sensors and
image_sensors.mspct contains spectral data for digital
cameras and image sensors.
Each spectrum object contains in addition to the spectral data, metadata stored as object attributes.
## List of 4
## $ model : chr " S10420-01 "
## $ supplier : chr "Hamamatsu"
## $ type : chr "CCD image sensor for spectromtres"
## $ entrance.optics: chr "none"
## - attr(*, "class")= chr [1:2] "sensor_properties" "list"
what_measured(camera.spct)## [1] "CCD image sensor for spectromtres S10420-01 from Hamamatsu"
how_measured(camera.spct)## [1] "Digitized from on-line plot from LUCID."## Data are approximate due to digitalization, smoothing and "wavelength-thinning".In addition to the objects containing the data itself, several character vectors of names of spectra are provide to facilitate the retrieval of subsets of spectra.
Accessing individual spectra
The response_spct member objects in collections
sensors.mspct and image_sensors.mspct can be
accessed through their names or through a numeric index. As the numeric
indexes are likely to change with updates to the package, their use is
discouraged. Names as character strings should be used instead. They can
also be retrieved with method names().
names(sensors.mspct)## [1] "ams_AS7263" "ams_AS7331"
## [3] "ams_AS7341" "ams_AS7343"
## [5] "ams_TSL254R" "ams_TSL257"
## [7] "ams_TSL2591" "AnalytikJena_UVX25"
## [9] "AnalytikJena_UVX31" "AnalytikJena_UVX36"
## [11] "apogee_s2_131_FR" "apogee_s2_131_R"
## [13] "apogee_sq_100X" "apogee_sq_500"
## [15] "apogee_sq_610" "apogee_su_200"
## [17] "Berger_UV_Biometer" "DeltaT_BF5"
## [19] "flat_e" "flat_q"
## [21] "Hamamatsu_G17191" "Hamamatsu_G17193"
## [23] "Hamamatsu_G6262" "Hamamatsu_R16571"
## [25] "Hamamatsu_S1226_BQ" "Irradian_DA211A_Cos"
## [27] "Irradian_DA211B2_Cos" "Irradian_DV211E_Cos"
## [29] "Irradian_DV211Q_Cos" "KIPP_CM21"
## [31] "KIPP_CUV_5" "KIPP_PQS1"
## [33] "KIPP_UVS_A" "KIPP_UVS_B"
## [35] "KIPP_UVS_E" "LICOR_LI_190"
## [37] "LICOR_LI_190R" "LICOR_LI_190SA"
## [39] "LICOR_LI_200" "LICOR_LI_210"
## [41] "LICOR_LI_210R" "LiteOn_LTR390"
## [43] "Osram_BPX65" "sglux_custom_green"
## [45] "sglux_custom_UVA1" "sglux_SG01D_A"
## [47] "sglux_SG01D_B" "sglux_SG01D_C"
## [49] "sglux_SG01L" "sglux_TOCON_blue4"
## [51] "Skye_SKE510" "Skye_SKL310"
## [53] "Skye_SKP210" "Skye_SKP215"
## [55] "Skye_SKR110_FR" "Skye_SKR110_R"
## [57] "Skye_SKS1110" "Skye_SKU421"
## [59] "Skye_SKU421a" "Skye_SKU430a"
## [61] "Skye_SKU440a" "SolarLight_501_Biometer_high_UVA"
## [63] "SolarLight_501_Biometer_low_UVA" "SolarLight_501_Biometer_typical"
## [65] "Solarmeter_SM60" "Specmeters_3415F"
## [67] "Thies_E1c" "Vishay_VEML6075"
## [69] "Vital_BW_20"
names(image_sensors.mspct)## [1] "Hamamatsu_S10420_01" "LUCID_TRT033S_WC_IMX992"
## [3] "LUCID_TRI071S_M_IMX428" "LUCID_ATX081S_UC_IMX487"
## [5] "LUCID_TRI071S_C_IMX428"We can use a character string as index to extract an individual
response_spct object.
sensors.mspct$LICOR_LI_190R## Object: response_spct [214 x 2]
## Wavelength range 380.76093-797.90271 nm, step 0.5007704-4.006164 nm
## Label: LICOR LI 190R light sensor
## Sensor: LI from LICOR.
## Spct: data in s.e.response normalized to 1 at 661.7 nm (max in 381-798 nm)
## Variables:
## w.length: Wavelength [nm]
## s.e.response: Spectral energy response [normalized]
## --
## # A tibble: 214 × 2
## w.length s.e.response
## <dbl> <dbl>
## 1 381. 0.00164
## 2 381. 0.00194
## 3 382. 0.00224
## 4 383. 0.00284
## 5 385. 0.00405
## 6 386. 0.00653
## 7 387. 0.00775
## 8 387. 0.00853
## 9 388. 0.00899
## 10 388. 0.0110
## # ℹ 204 more rows
sensors.mspct[["LICOR_LI_190R"]]## Object: response_spct [214 x 2]
## Wavelength range 380.76093-797.90271 nm, step 0.5007704-4.006164 nm
## Label: LICOR LI 190R light sensor
## Sensor: LI from LICOR.
## Spct: data in s.e.response normalized to 1 at 661.7 nm (max in 381-798 nm)
## Variables:
## w.length: Wavelength [nm]
## s.e.response: Spectral energy response [normalized]
## --
## # A tibble: 214 × 2
## w.length s.e.response
## <dbl> <dbl>
## 1 381. 0.00164
## 2 381. 0.00194
## 3 382. 0.00224
## 4 383. 0.00284
## 5 385. 0.00405
## 6 386. 0.00653
## 7 387. 0.00775
## 8 387. 0.00853
## 9 388. 0.00899
## 10 388. 0.0110
## # ℹ 204 more rowsBe aware that according to R’s rules, using single square brackets
will return a response_mspct object possibly of length one.
This statement is not equivalent to the one in the chunk immediately
above.
sensors.mspct["LICOR_LI_190R"]## Object: response_mspct [1 x 1]
## --- Member: LICOR_LI_190R ---
## Object: response_spct [214 x 2]
## Wavelength range 380.76093-797.90271 nm, step 0.5007704-4.006164 nm
## Label: LICOR LI 190R light sensor
## Sensor: LI from LICOR.
## Spct: data in s.e.response normalized to 1 at 661.7 nm (max in 381-798 nm)
## Variables:
## w.length: Wavelength [nm]
## s.e.response: Spectral energy response [normalized]
## --
## # A tibble: 214 × 2
## w.length s.e.response
## <dbl> <dbl>
## 1 381. 0.00164
## 2 381. 0.00194
## 3 382. 0.00224
## 4 383. 0.00284
## 5 385. 0.00405
## 6 386. 0.00653
## 7 387. 0.00775
## 8 387. 0.00853
## 9 388. 0.00899
## 10 388. 0.0110
## # ℹ 204 more rows
##
## --- END ---Accessing subsets of spectra
We can subset sensors.mspct object by indexing with
vectors of character strings. The package provides some predefined ones,
and users can easily define their own, either as constants or through
computation. Here we use a vector defined by the package.
sensors.mspct[berger_sensors]## Object: response_mspct [1 x 1]
## --- Member: Berger_UV_Biometer ---
## Object: response_spct [21 x 2]
## Wavelength range 279.872-378.458 nm, step 3.311-5.284 nm
## Label: Berger UV Biometer light sensor
## Sensor: UV from Berger.
## Spct: data in s.e.response normalized to 1 at 290.2 nm (max in 280-378 nm)
## Variables:
## w.length: Wavelength [nm]
## s.e.response: Spectral energy response [normalized]
## --
## # A tibble: 21 × 2
## w.length s.e.response
## <dbl> <dbl>
## 1 280. 0.747
## 2 285. 0.912
## 3 290. 1
## 4 295. 0.985
## 5 300. 0.770
## 6 305. 0.471
## 7 310. 0.215
## 8 315. 0.0782
## 9 320. 0.0263
## 10 325. 0.00845
## # ℹ 11 more rows
##
## --- END ---More generally one can search for matching names within the collection of spectra.
## Object: response_mspct [1 x 1]
## --- Member: Berger_UV_Biometer ---
## Object: response_spct [21 x 2]
## Wavelength range 279.872-378.458 nm, step 3.311-5.284 nm
## Label: Berger UV Biometer light sensor
## Sensor: UV from Berger.
## Spct: data in s.e.response normalized to 1 at 290.2 nm (max in 280-378 nm)
## Variables:
## w.length: Wavelength [nm]
## s.e.response: Spectral energy response [normalized]
## --
## # A tibble: 21 × 2
## w.length s.e.response
## <dbl> <dbl>
## 1 280. 0.747
## 2 285. 0.912
## 3 290. 1
## 4 295. 0.985
## 5 300. 0.770
## 6 305. 0.471
## 7 310. 0.215
## 8 315. 0.0782
## 9 320. 0.0263
## 10 325. 0.00845
## # ℹ 11 more rows
##
## --- END ---Set algebra operations can be used with the indexing vectors as each vector describes a single property: color, brand, type, etc.
sensors.mspct[intersect(licor_sensors, par_sensors)]## Object: response_mspct [1 x 1]
## --- Member: LICOR_LI_190 ---
## Object: response_spct [287 x 2]
## Wavelength range 381.85375-798.83668 nm, step 0.5005798-4.004638 nm
## Label: LICOR LI 190 light sensor
## Sensor: LI from LICOR.
## Spct: data in s.e.response normalized to 1 at 676.9 nm (max in 382-799 nm)
## Variables:
## w.length: Wavelength [nm]
## s.e.response: Spectral energy response [normalized]
## --
## # A tibble: 287 × 2
## w.length s.e.response
## <dbl> <dbl>
## 1 382. 0.0272
## 2 382. 0.0291
## 3 383. 0.0310
## 4 384. 0.0349
## 5 386. 0.0425
## 6 389. 0.0559
## 7 389. 0.0590
## 8 390. 0.0613
## 9 390. 0.0650
## 10 391. 0.0689
## # ℹ 277 more rows
##
## --- END ---Calculating summaries from the normalized data
The spectra are normalized, and consequently, several summaries expressed in absolute units are undefined, and trigger errors. Summaries like ratios which are not affected by normalization are allowed and valid. The data have been normalized as the measuring conditions used are not all the same, and in many cases not well characterized (e.g. distance to nearby reflecting walls, or exact alignment of the spectrometer input optics with respect to light sources).
What we will do in this section is to rescale the spectral data so that after conversion a given target value for a summary quantity will be true. As an example, we will rescale one spectrum so that it yields a response of 100 mol-1 m2for the range 400 to 700 nm.
my.spct <- fscale(sensors.mspct$LICOR_LI_190R,
range = PhR(),
q_response,
target = 100,
set.scaled = FALSE)
q_response(my.spct, PhR())## R[/q]_PhR
## 100
## attr(,"time.unit")
## [1] "second"
## attr(,"radiation.unit")
## [1] "total photon response"
q_response(my.spct, UVA())## R[/q]_]UVA.ISO
## 0.3073047
## attr(,"time.unit")
## [1] "second"
## attr(,"radiation.unit")
## [1] "total photon response"If we want to treat the rescaled spectral data, as if they were true
readings with no scaling we can reset the attribute with method
setScaled(). With method getScaled() we can
test if a spectrum has been scaled.
getScaled(my.spct)## [1] FALSEIf for some obscure reason we want to simply “pretend” that the spectral data have not been normalized, we can permanently override the attribute on a copy of the data. Most of the time this is a very bad idea!
my2nd.spct <- sensors.mspct$LICOR_LI_190
setNormalized(my2nd.spct)
q_response(my2nd.spct)## R[/q]_Total
## 49403342
## attr(,"time.unit")
## [1] "second"
## attr(,"radiation.unit")
## [1] "total photon response"Plotting the spectra
Using autoplot() methods for spectra defined in package
‘ggspectra’ annotated plotting is automatic. The defaults can be easily
changed, please see the documentation in package ‘ggspectra’.
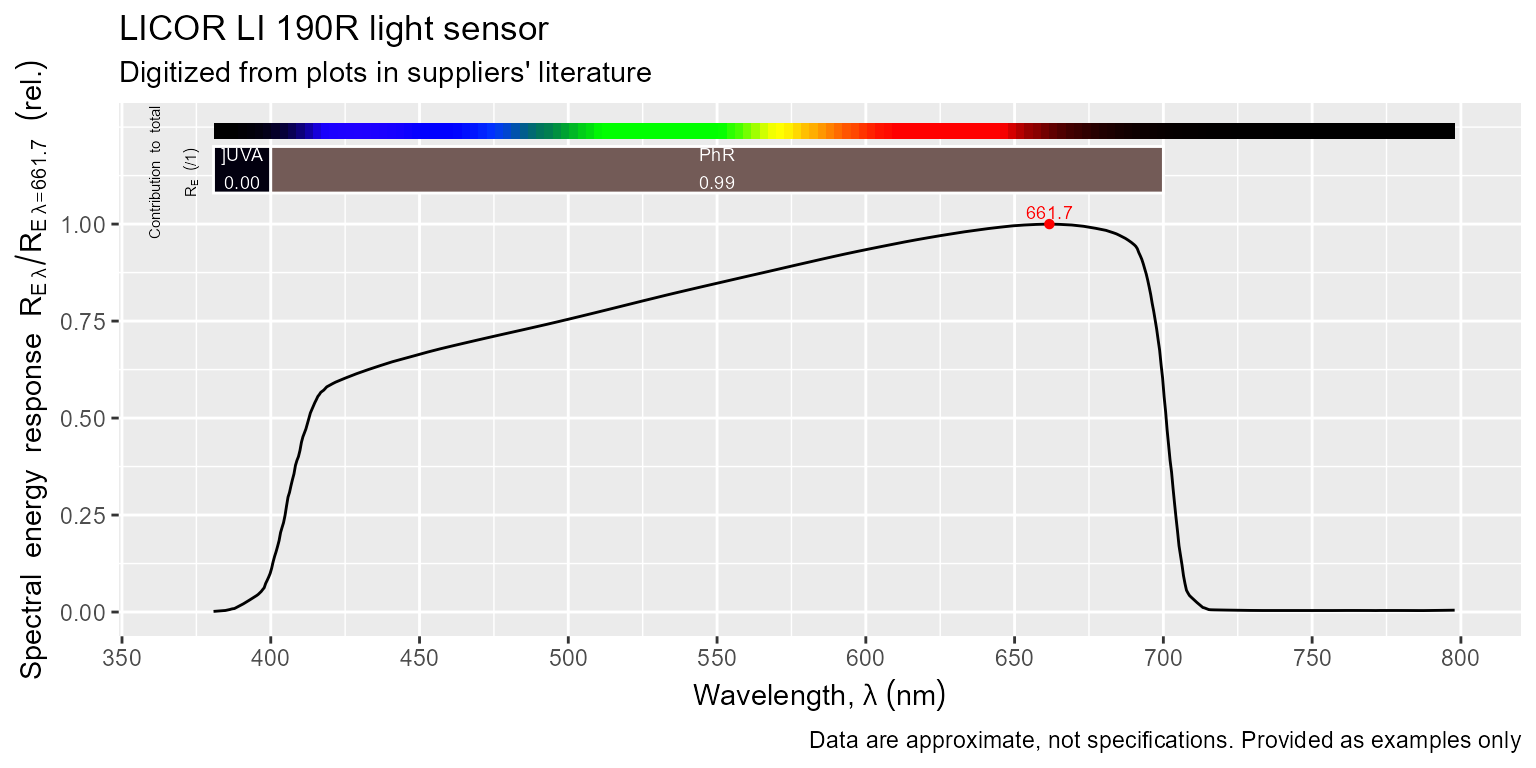
Using the ggplot() method for spectra from package
‘ggspectra’ plus geometries and statistics from
package ‘ggplot2’ we gain additional control on the design.
ggplot(sensors.mspct$LICOR_LI_190R, unit.out = "photon") +
geom_hline(yintercept = 1, colour = "red") +
geom_hline(yintercept = c(0.9, 1.1), colour = "red", linetype = "dotted") +
geom_line(linetype = "dashed") +
scale_y_s.q.response_continuous(breaks = c(0, 0.5, 1)) +
scale_x_wl_continuous() +
theme_classic()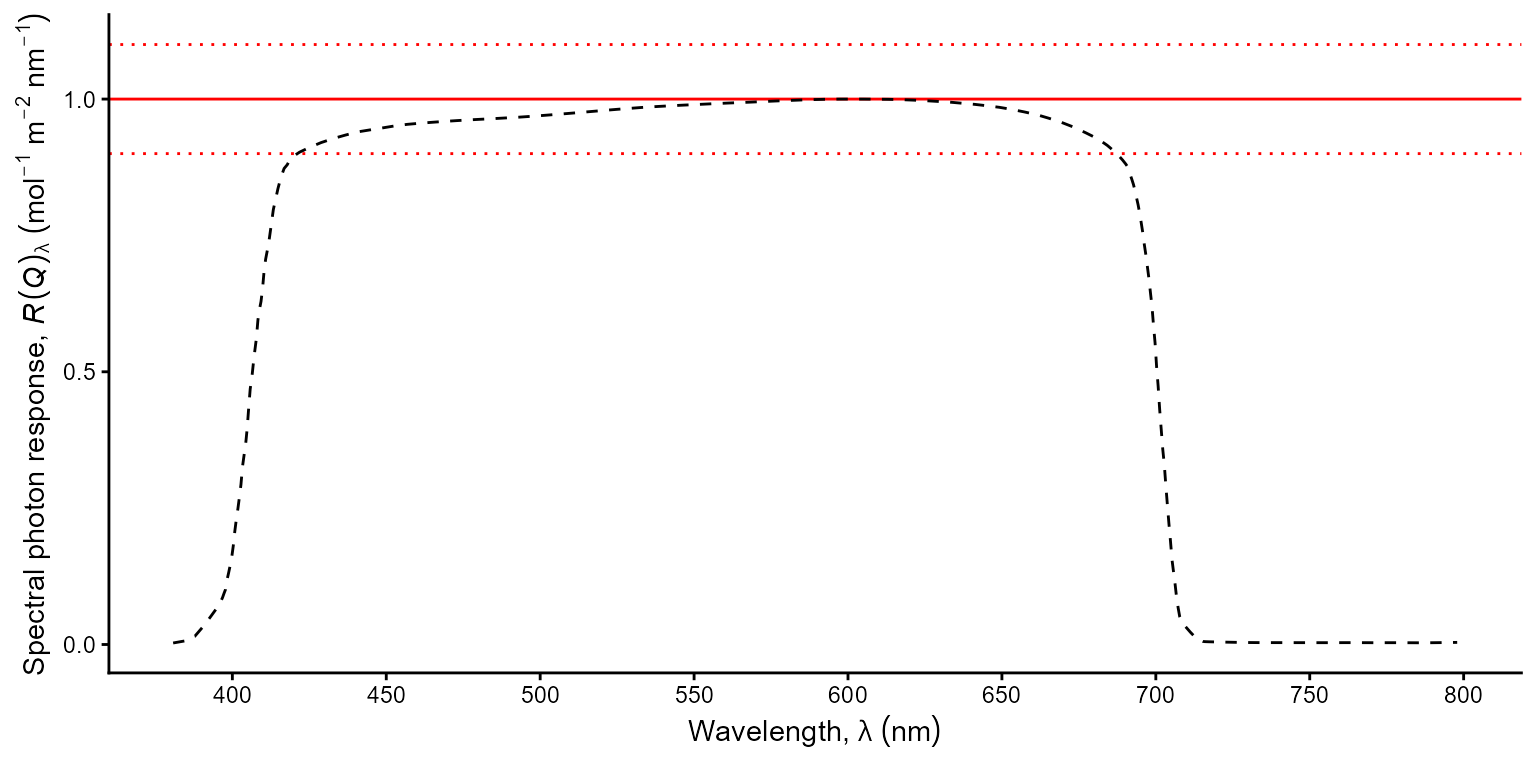
Angular response
ggplot(diffusers.lst$bentham_D7, aes(angle.deg, response)) +
geom_line() +
geom_line(aes(y = cos(angle.deg * pi / 180)), linetype = "dotted", color = "red") +
scale_x_continuous(name = "Angle (degrees)", breaks = c(-90, -60, -30, 0, 30, 60, 90)) +
scale_y_continuous(name = "Response (/1)") +
theme_classic()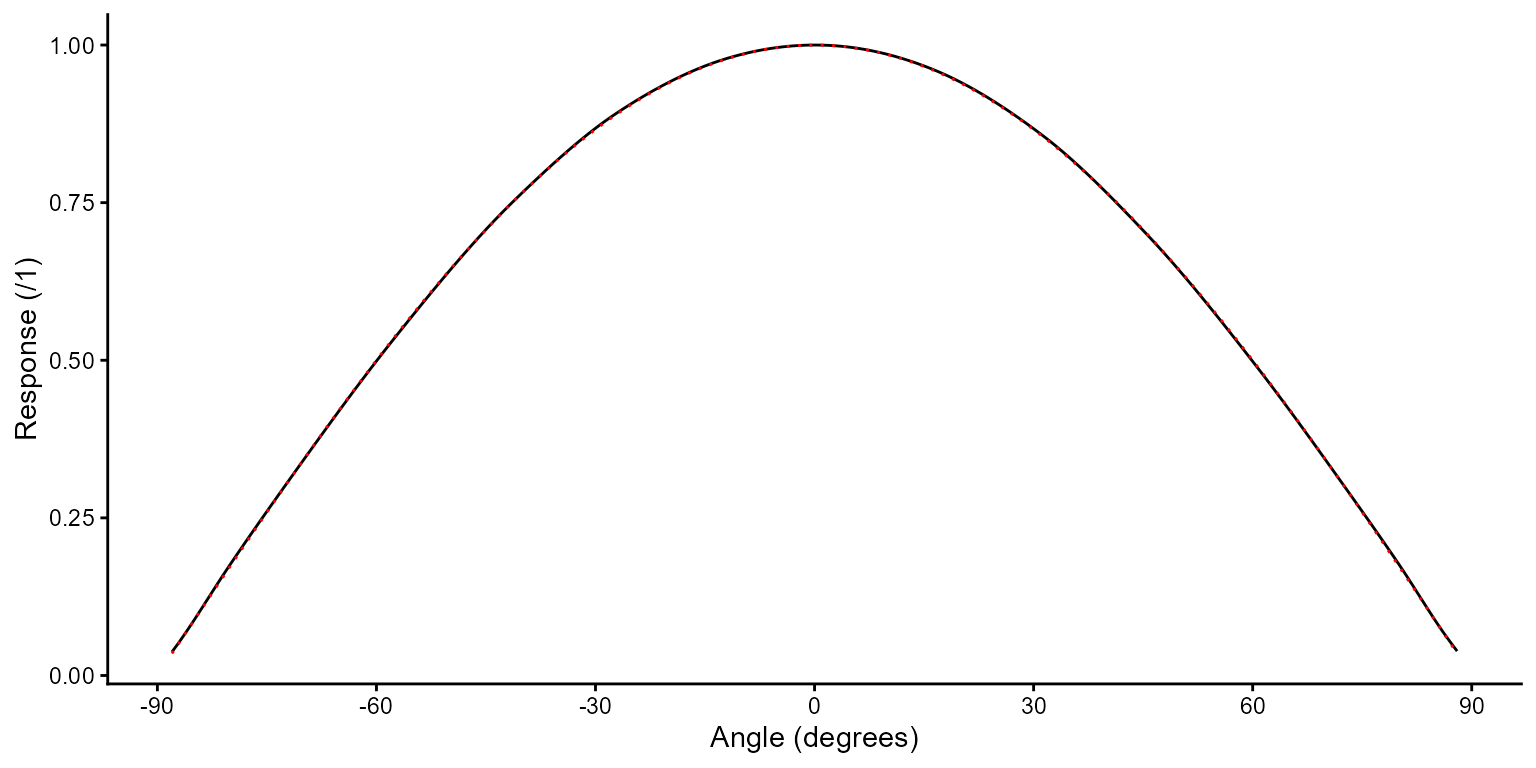
angular_response(45, "cosine")## [1] 0.7071068
angular_response(45, "dome")## [1] 0.8535534
angular_response(45, "sphere")## [1] 1Using the data in other contexts
As response_mspct is a class derived from
list, and response_spct is derived from
tibble::tible which is a (mostly) compatible
reimplementation of data.frame the data can be used very
easily with any R function.
head(as.data.frame(sensors.mspct$LICOR_LI_190R))## w.length s.e.response
## 1 380.7609 0.001644993
## 2 381.2617 0.001943170
## 3 381.7625 0.002242125
## 4 382.7640 0.002842367
## 5 384.7671 0.004052183
## 6 386.2694 0.006525098Of course attach and with also work as
expected.
## Object: response_spct [6 x 2]
## Wavelength range 380.76093-386.2694 nm, step 0.5007704-2.003082 nm
## Label: LICOR LI 190R light sensor
## Sensor: LI from LICOR.
## Spct: data in s.e.response normalized to 1 at 661.7 nm (max in 381-798 nm)
## Variables:
## w.length: Wavelength [nm]
## s.e.response: Spectral energy response [normalized]
## --
## # A tibble: 6 × 2
## w.length s.e.response
## <dbl> <dbl>
## 1 381. 0.00164
## 2 381. 0.00194
## 3 382. 0.00224
## 4 383. 0.00284
## 5 385. 0.00405
## 6 386. 0.00653
detach(sensors.mspct)## [1] 798.8367
detach(sensors.mspct)## [1] 381.8537 798.8367