
Fitted model prediction and annotations
Source:R/stat-poly-eq.R, R/stat-poly-line.R
stat_poly_eq.RdStatistics stat_poly_line and stat_poly_eq() fit a
model, by default with stats::lm(), but alternatively using other
model fit functions. While stat_poly_line adds a prediction line and
band, stat_poly_eq() adds textual labels to a plot.
Usage
stat_poly_eq(
mapping = NULL,
data = NULL,
geom = "text_npc",
position = "identity",
...,
orientation = NA,
formula = NULL,
method = "lm",
method.args = list(),
n.min = 2L,
fit.seed = NA,
eq.with.lhs = TRUE,
eq.x.rhs = NULL,
small.r = getOption("ggpmisc.small.r", default = FALSE),
small.p = getOption("ggpmisc.small.p", default = FALSE),
CI.brackets = c("[", "]"),
rsquared.conf.level = 0.95,
coef.digits = 3,
coef.keep.zeros = TRUE,
decreasing = getOption("ggpmisc.decreasing.poly.eq", FALSE),
rr.digits = 2,
f.digits = 3,
p.digits = 3,
label.x = "left",
label.y = "top",
hstep = 0,
vstep = NULL,
output.type = NULL,
na.rm = FALSE,
parse = NULL,
show.legend = FALSE,
inherit.aes = TRUE
)
stat_poly_line(
mapping = NULL,
data = NULL,
geom = "smooth",
position = "identity",
...,
orientation = NA,
method = "lm",
formula = NULL,
se = NULL,
fit.seed = NA,
fm.values = FALSE,
n = 80,
fullrange = FALSE,
level = 0.95,
method.args = list(),
n.min = 2L,
na.rm = FALSE,
show.legend = NA,
inherit.aes = TRUE
)Arguments
- mapping
The aesthetic mapping, usually constructed with
aes(). Only needs to be set at the layer level if you are overriding the plot defaults.- data
A layer specific dataset, only needed if you want to override the plot defaults.
- geom
The geometric object to use display the data
- position
The position adjustment to use for overlapping points on this layer.
- ...
other arguments passed on to
layer. This can include aesthetics whose values you want to set, not map. Seelayerfor more details.- orientation
character Either "x" or "y" controlling the default for
formula. The letter indicates the aesthetic considered the explanatory variable in the model fit.- formula
a formula object. Using aesthetic names
xandyinstead of original variable names.- method
function or character If character, "lm", "rlm", "lmrob", "lts", "gls", "ma", "sma", "segreg", "rq" or the name of a model fit function are accepted, possibly followed by the fit function's
methodargument separated by a colon (e.g."rlm:M"). If a function is different tolm(),rlm(),ltsReg(),gls(),ma,sma, it must have formal parameters namedformula,data, andweights. See Details.- method.args
named list with additional arguments. Not
dataorweightswhich are always passed through aesthetic mappings.- n.min
integer Minimum number of distinct values in the explanatory variable (on the rhs of formula) for fitting to the attempted.
- fit.seed
RNG seed argument passed to
set.seed(). Defaults toNA, indicating thatset.seed()should not be called.- eq.with.lhs
If
characterthe string is pasted to the front of the equation label before parsing or alogical(see note).- eq.x.rhs
characterthis string will be used as replacement for"x"in the model equation when generating the label before parsing it.- small.r, small.p
logical Flags to switch use of lower case r and p for coefficient of determination and p-value.
- CI.brackets
character vector of length 2. The opening and closing brackets used for the CI label.
- rsquared.conf.level
numeric Confidence level for the returned confidence interval. Set to NA to skip CI computation.
- coef.digits, f.digits
integer Number of significant digits to use for the fitted coefficients and F-value.
- coef.keep.zeros
logical Keep or drop trailing zeros when formatting the fitted coefficients and F-value.
- decreasing
logical It specifies the order of the terms in the returned character string; in increasing (default) or decreasing powers.
- rr.digits, p.digits
integer Number of digits after the decimal point to use for \(R^2\) and P-value in labels. If
Inf, use exponential notation with three decimal places.- label.x, label.y
numericwith range 0..1 "normalized parent coordinates" (npc units) or character if usinggeom_text_npc()orgeom_label_npc(). If usinggeom_text()orgeom_label()numeric in native data units. If too short they will be recycled.- hstep, vstep
numeric in npc units, the horizontal and vertical step used between labels for different groups.
- output.type
character One of "expression", "LaTeX", "text", "markdown" or "numeric".
- na.rm
a logical indicating whether NA values should be stripped before the computation proceeds.
- parse
logical Passed to the geom. If
TRUE, the labels will be parsed into expressions and displayed as described inplotmath. Default isTRUEifoutput.type = "expression"andFALSEotherwise.- show.legend
logical. Should this layer be included in the legends?
NA, the default, includes if any aesthetics are mapped.FALSEnever includes, andTRUEalways includes.- inherit.aes
If
FALSE, overrides the default aesthetics, rather than combining with them. This is most useful for helper functions that define both data and aesthetics and shouldn't inherit behaviour from the default plot specification, e.g.borders.- se
Display confidence interval around smooth? (`TRUE` by default only for fits with
lm()andrlm(), see `level` to control.)- fm.values
logical Add metadata and parameter estimates extracted from the fitted model object;
FALSEby default.- n
Number of points at which to predict with the fitted model.
- fullrange
Should the fit span the full range of the plot, or just the range of the data group used in each fit?
- level
Level of confidence interval to use (0.95 by default).
Value
stat_poly_eq() returns a data frame, with a single row per
group and columns as described below. stat_poly_line() returns a
data frame, with n rows per group and columns as described below. In
cases when the number of observations is less than n.min or when the
model fit function returns a single NA or NULL, a data frame
with no rows or columns (built by data.frame()) is returned, and
silently rendered as an empty/invisible plot layer.
When a predict() method is not available for the fitted model class,
the value returned by calling fitted(), if available, is replaces it
and the returned data frame with as many rows as observations, instead of
n rows, is returned with a message.
Details
Statistics stat_poly_line() and stat_poly_eq()
fit a model consistently, but return different values.
stat_poly_line() plots a prediction line and band, similarly to
stat_smooth()
but has different defaults and supports a different set of model fit
functions.
stat_poly_eq() adds textual labels for
\(R^2\), adjusted \(R^2\), the fitted model equation, \(P\), and
other parameters from a fitted model to a plot.
Lack of methods or explicit support for extraction of individual parameters
results in the affected estimates and corresponding labels being set to
NA. Similarly, confidence bands for the prediction line are not
plotted in some cases, while in the case of MA and SMA models, the band
only displays the uncertainty of the slope rather than for both slope plus
intercept. While strings for \(R^2\), adjusted \(R^2\), \(F\), and
\(P\) annotations are returned for all valid linear models and many other
types of fitted models, an automatically constructed character string for
the fitted model equation is returned only for polynomials (see below).
However, when not generated automatically, the equation can still be
assembled by the user within the call to aes(). A
label for the confidence interval of \(R^2\), based on values computed
with function ci_rsquared() from package 'confintr'
is returned when possible.
When possible, i.e., nearly always, the formula used to build the
equation label is extracted from the returned fitted model object. Most
fitted model objects follow the example of lm() and include the
formula for the model that has been fitted. Thus, this model formula can
safely differ from the argument passed to parameter formula in the
call to stat_poly_eq().
The stats are designed to support user-defined methods that
implement any or all of method selection, model formula
selection, dynamically adjusted method.args and conditional skipping
of labelling on a by group basis.
The minimum number of observations with distinct values in the explanatory
variable can be set through parameter n.min. The default n.min
= 2L is the smallest suitable for method "lm" but too small for
method "rlm" for which n.min = 3L is needed. Anyway, model
fits with very few observations are of little interest and using larger
values of n.min than the default is wise.
As some model fitting approaches depend on the RNG (pseudo-Random Number
Generator), when fit.seed is not NA it is used as argument in
a call to set.seed() immediately ahead of model
fitting, i.e., once for each group of observations.
Singularity, convergence, etc., are handled by the model fit functions.
With method "lm", singularity results in terms being dropped with a
message if more numerous than can be fitted with a singular (exact) fit. In
this case and if the model results in a perfect fit due to low number of
observation, estimates for various parameters are NaN or NA.
With methods other than "lm", the model fit functions simply fail in
case of singularity, e.g., singular fits are not implemented in
"rlm".
Model formula and model fitting
A ggplot statistic receives as data a data frame that is not the one
passed as argument by the user, but instead a data frame with the variables
mapped to aesthetics. In stat_poly_eq() the compute function is
applied by group, each call "seeing" the subset of data for an
individual group. As supported models are for regression lines,
variables mapped to x and y should both be continuous, i.e.,
numeric or date time and model formulas defined using x and y
as variable names.
The interpretation of the argument passed to formula is enhanced
compared to stat_smooth(). Formulas with x as explanatory
variable work as in stat_smooth() but formulas with y as
explanatory variable are also accepted. orientation is set
automatically based on which explanatory variable appears in the formula.
Spline-based smoothers are only partially supported.
Model equation label
By default the equation label uses as symbols the names of the aesthetics,
x and y. However, "x" and "y" can be
substituted by providing a replacement character string for the
right-hand-side and left-hand-side through eq.x.rhs and
eq.with.lhs, respectively. For backward compatibility a logical is
also accepted as argument for eq.with.lhs, with FALSE
suppressing the left-hand-side.
If the model formula includes a transformation of the explanatory
variable in its right-hand-side (rhs), a matching argument should be passed
to parameter eq.x.rhs as its default value would result in an
equation label that does not reflect the applied transformation. In most
cases, a transformation should not be applied within the left hand side
(lhs) of the model formula, but instead in the mapping of the response
variable within aes. In this case it may be necessary to also pass a
matching argument to parameter eq.with.lhs.
Parameter orientation is redundant as the orientation can be set
by the formula but is included for consistency with
ggplot2::stat_smooth().
Position of labels
When data are grouped by mapping a factor to an aesthetic, e.g.,
colour, shape and/or linetype the model is fitted
separately to each group, and for each group a whole set of labels is
generated. If the argument passed to label.y is a vector of length
1, this value determines the position of the equation and/or other labels
for the first group, and the positions of the labels for the remaining
groups are generated by adding vspace based on the group number.
If the argument passed to label.y is a vector of length > 1, it is
used unchanged, possibly extended by recycling, ignoring vstep.
If the labels are rotated by 90 degrees then the automatic stepping is
best based on hstep with vstep = 0. Similarly as described
above, if label.x is a vector of length > 1, it is
used unchanged, possibly extended by recycling, ignoring hstep.
When using facets and with a grouping that does not repeat in each panel,
the automatic positioning in most cases will not be the desired one. Manual
positioning using a vector of length > 1 for label.x and/or
label.y is the currently available workaround.
Model fit methods supported
Several model fit functions are supported explicitly (see tables), and some
of their differences smoothed out. Compatibility is checked late, based on
the class of the returned fitted model object. This makes it possible to
use wrapper functions that do model selection or other adjustments to the
fit procedure on a per panel or per group basis. Moreover, if the value
returned as model fit object is NULL or NA, plotting is
skipped on a per group within panel basis.
In the case of fitted model objects of classes not explicitly supported, an attempt is made to find the usual accessors and/or fitted object members, and if found, either complete or partial support is frequently achieved. In this case a message is issued encouraging users to check the validity of the values extracted as the structure of fitted model objects belonging to different classes and the values returned by their accessors can vary, potentially resulting in decoding errors leading to the return of wrong values for estimates.
The argument to parameter method can be either the name of a
function object, possibly using double colon notation in case its package
is not attached, or a character string matching the function name for
functions in the search path. This approach makes it possible to support
model fit functions that are not dependencies of 'ggpmisc'. Either by
attaching the package where the function is defined and passing it by name
or as string, or using double colon notation when passing the name of the
function.
User-defined functions can be passed as argument to parameter method
as long as they have parameters formula, data subset
and possibly weights. Additional arguments can be passed to any
method as a named list through parameter method.args. As in
stat_smooth() prior weights are
passed to the model fit functions' weights (plural!) parameter by
mapping a numeric variable to plot aesthetic weight (singular!).
Tables 1 lists natively supported model fit functions, with the caveat that only some 'broom' methods' specializations have been actually tested with statistics from 'ggpmisc'. In addition, the statistics based on 'broom' methods require the user to tailor their behaviour by passing additional arguments in the call and occasionally some detective work to find out the names of variables in the returned data frame as these names are set by methods from 'broom'.
Table 1. Model fit methods supported by the different statistics available in package 'ggpmisc'. Column \(f\) indicates whether computations are done by group (G) or by plot panel (P).
| Statistic | \(f\) | Supported model fit methods |
stat_poly_line() | G | "lm", "rlm", "lts", "sma", "ma", "gls", others with methods predict() or fitted() |
stat_poly_eq() | G | "lm", "rlm", "lts", "sma", "ma", "gls", others with needed accesors |
stat_quant_line() | G | "rq", "rqss" |
stat_quant_band() | G | "rq", "rqss" |
stat_quant_eq() | G | "rq", "rqss" |
stat_ma_line() | G | "SMA", "MA", "RMA", "OLS" |
stat_ma_eq() | G | "SMA", "MA", "RMA", "OLS" |
stat_fit_residuals() | G | "lm", "rlm", "lts", "sma", "ma", "gls", "rq", "rqss" others with method residuals() |
stat_fit_fitted() | G | "lm", "rlm", "lts", "gls", "rq", "rqss" others with method fitted() |
stat_fit_deviations() | G | "lm", "rlm", "lts", "gls", "rq", "rqss" others with methods fitted() and weights() |
stat_fit_augment() | G | any with 'broom' method augment() |
stat_fit_glance() | G | any with 'broom' method glance() |
stat_fit_tidy() | G | any with 'broom' method tidy() |
stat_fit_tb() | P | any with 'broom' method tidy() |
The single colon notation is based on parsing
the name and is available when passing the name of the fit method as a
character string. In a string such as "head:tail" the "head" gives the name
of the model fit function and the "tail" gives the argument to pass it's
method parameter. This is only a convenience, as method.args
can be also used. In some methods, i.e., splines, the default
formula = y ~ x needs to be overridden by the user.
Table 2 lists the correspondence of pre-defined method names to model fit method functions. As mentioned above, these are only a subset of the model fit methods that are expected to work. When using these names there is no need for users to attach additional packages but the packages must be available (installed).
Table 2. Available predefined method names, the model fit functions
they call, the packages where the functions reside, the class of the
returned fitted model object and the arguments that can be
passed to their method parameter using single colon notation.
| Predefined method names | Model fit methods | R package | Object class |
| "lm", "lm:qr" | lm() | 'stats' | "lm" |
| "rlm", "rlm:M", "rlm:MM" | rlm() | 'MASS' | "rlm" ("lm") |
| "lts", "ltsReg" | ltsReg() | 'robustbase' | "lts" |
| "ma", "sma", "sma:SMA", "sma:MA", "sma:OLS" | sma() | 'smatr' | "ma" or "sma" |
| "gls", "gls:REML", "gls:ML" | gls() | 'nlme' | "gls" |
| "rq", "rq:sfn", "rq:sfnc", "rq:lasso" | rq() | 'quantreg' | "rq" |
| "rqss", "rqss:sfn", "rqss:sfnc", "rqss:lasso" | rqss() | 'quantreg' | "rqss" |
| "SMA", "MA", "RMA", "OLS" | lmodel2() | 'lmodel2' | ("list") |
Variables returned by `stat_poly_line()`
Some of the variables depend on the orientation.
- y or x
predicted value
- ymin or xmin
lower confidence limit around the fitted line
- ymax or xmax
upper confidence limit around the fitted line
- se
standard error
If fm.values = TRUE is passed then columns based on the summary of
the model fit are added, with the same value in each row within a group.
This is wasteful and disabled by default, but provides a simple and robust
approach to achieve effects like colouring or hiding of the model fit line
based on \(P\), \(R^2\), \(R_{adj}^2\) or the number of
observations in a fit.
Variables returned by stat_poly_eq()
For all output.type arguments the following values are returned.
- x,npcx
x position
- y,npcy
y position
- coefs
fitted coefficients, named numeric vector as a list member
- r.squared, rr.confint.level, rr.confint.low, rr.confint.high, adj.r.squared, f.value, f.df1, f.df2, p.value, AIC, BIC, n, knots, knots.se
numeric values, from the model fit object
- grp.label
Set according to mapping in
aes.- knots
list containing a numeric vector of knot or "psi" x-value for linear splines
- fm.method
name of method used, character
- fm.class
most derived class or the fitted model object, character
- fm.formula.chr
formatted model formula, character
If output.type is not "numeric" the returned tibble contains in
addition to those above the columns listed below, each containing a single
character string. The markup used depends on the value of output.type.
- eq.label
equation for the fitted polynomial as a character string to be parsed or
NA- rr.label
\(R^2\) of the fitted model as a character string to be parsed
- adj.rr.label
Adjusted \(R^2\) of the fitted model as a character string to be parsed
- rr.confint.label
Confidence interval for \(R^2\) of the fitted model as a character string to be parsed
- f.value.label
F value and degrees of freedom for the fitted model as a whole.
- p.value.label
P-value for the F-value above.
- AIC.label
AIC for the fitted model.
- BIC.label
BIC for the fitted model.
- n.label
Number of observations used in the fit.
- knots.label
The knots or change points in segmented regression.
- grp.label
Set according to mapping in
aes.- method.label
Set according
methodused.
If output.type is "numeric" the returned tibble contains columns
listed below in addition to the base ones. If the model fit function used
does not return a value, the variable is set to NA_real_.
- coef.ls
list containing the "coefficients" matrix from the summary of the fit object
- b_0.constant
TRUE is polynomial is forced through the origin
- b_i
One or more columns with the coefficient estimates
To explore the computed values returned for a given input we suggest the use
of geom_debug() as shown in the last examples below.
Output types
The formatting of character strings to be displayed in plots are marked as mathematical equations. Depending on the geom used, the mark-up needs to be encoded differently, or in some cases mark-up not applied.
"expression"The labels are encoded as character strings to be parsed into R's plotmath expressions.
"LaTeX", "TeX", "tikz", "latex"The labels are encoded as 'LaTeX' maths equations, without the "fences" for switching in math mode.
"latex.eqn"Same as
"latex"but enclosed in single$, i.e., as in-line maths."latex.deqn"Same as
"latex"but enclosed in double$$, i.e., as display maths."markdown"The labels are encoded as character strings using markdown syntax, with some embedded HTML.
"marquee"The labels are encoded as character strings using markdown syntax, with 'marquee' supported spans.
"text"The labels are plain ASCII character strings.
"numeric"No labels are generated. This value is accepted by the statistics, but not by the label formatting functions.
NULLThe value used,
expression,latex.eqnormarkupdepends on the argument passed togeom.
If geom = "latex" (package 'xdvir') the output type used is
"latex.eqn". If geom = "richtext" (package 'ggtext') or
geom = "textbox" (package 'ggtext') the output type used is
"markdown". If geom = "marquee" (package 'marquee') the output
type used is "marquee". For all other values of geom the default
is "expression" unless the user passes an argument. Invalid values as
argument trigger an Error.
References
Originally written as an answer to question 7549694 at Stackoverflow but enhanced based on suggestions from several users and my own needs.
See also
Consult the documentation of the model fit functions used
for the details and additional arguments that can be passed to
them by name through parameter method.args.
Please, see the articles in online-only documentation for additional use examples and guidance.
Other 'ggpmisc' statistics for model fits:
stat_distrmix_eq(),
stat_fit_deviations(),
stat_fit_glance(),
stat_fit_tb(),
stat_fit_tidy(),
stat_ma_eq(),
stat_quant_band()
Aesthetics
stat_poly_eq() understands the following aesthetics. Required aesthetics are displayed in bold and defaults are displayed for optional aesthetics:
| • | x | |
| • | y | |
| • | group | → inferred |
| • | grp.label | |
| • | hjust | → "inward" |
| • | label | → after_stat(rr.label) |
| • | npcx | → after_stat(npcx) |
| • | npcy | → after_stat(npcy) |
| • | vjust | → "inward" |
| • | weight | → 1 |
stat_poly_line() understands the following aesthetics. Required aesthetics are displayed in bold and defaults are displayed for optional aesthetics:
| • | x | |
| • | y | |
| • | group | → inferred |
Learn more about setting these aesthetics in vignette("ggplot2-specs").
Examples
# generate artificial data
set.seed(4321)
x <- 1:100
y <- (x + x^2 + x^3) + rnorm(length(x), mean = 0, sd = mean(x^3) / 4)
y <- y / max(y)
my.data <- data.frame(x = x, y = y,
group = c("A", "B"),
y2 = y * c(1, 2) + c(0, 0.1),
w = sqrt(x))
# give a name to a formula
formula <- y ~ poly(x, 3, raw = TRUE)
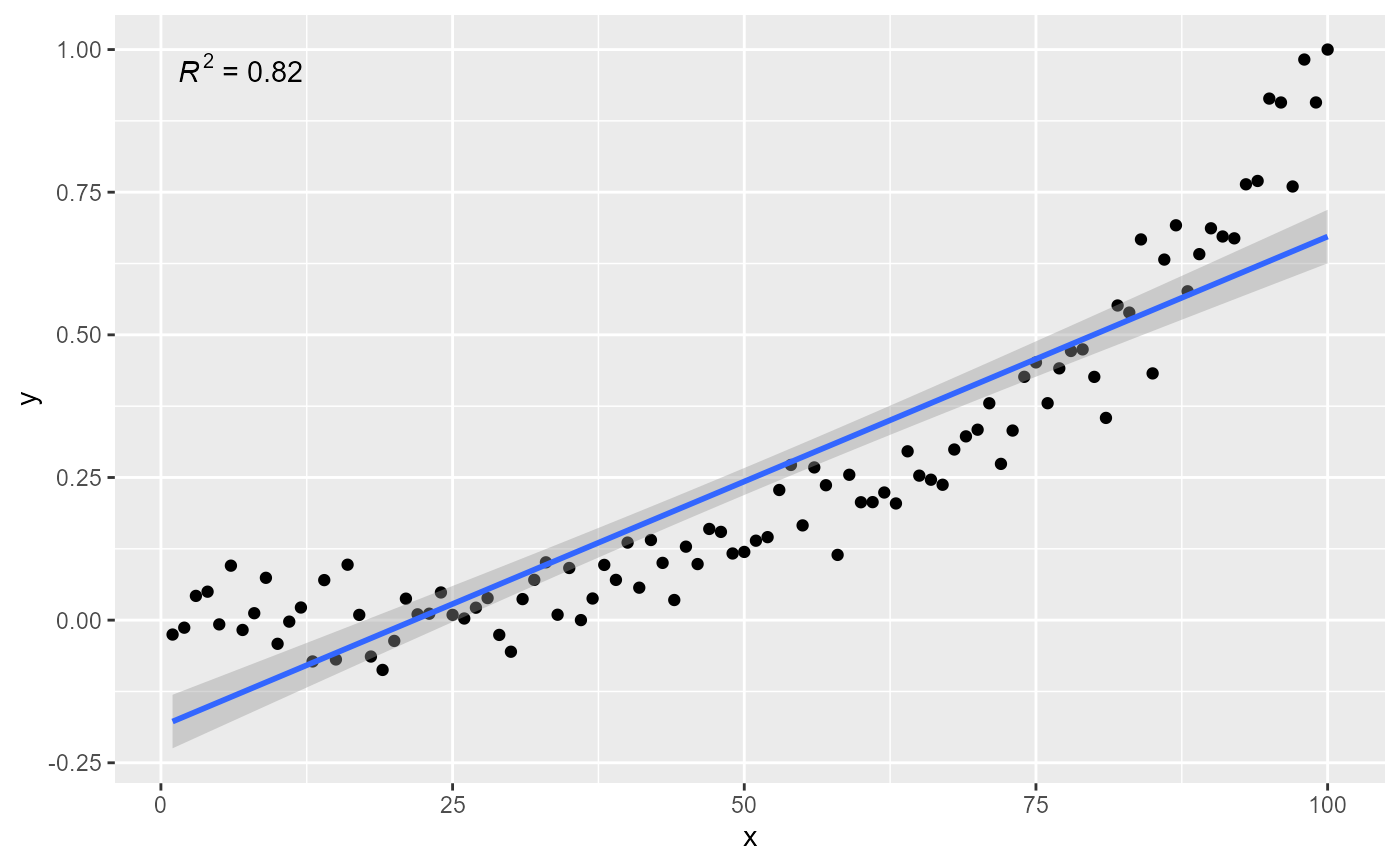
# using defaults
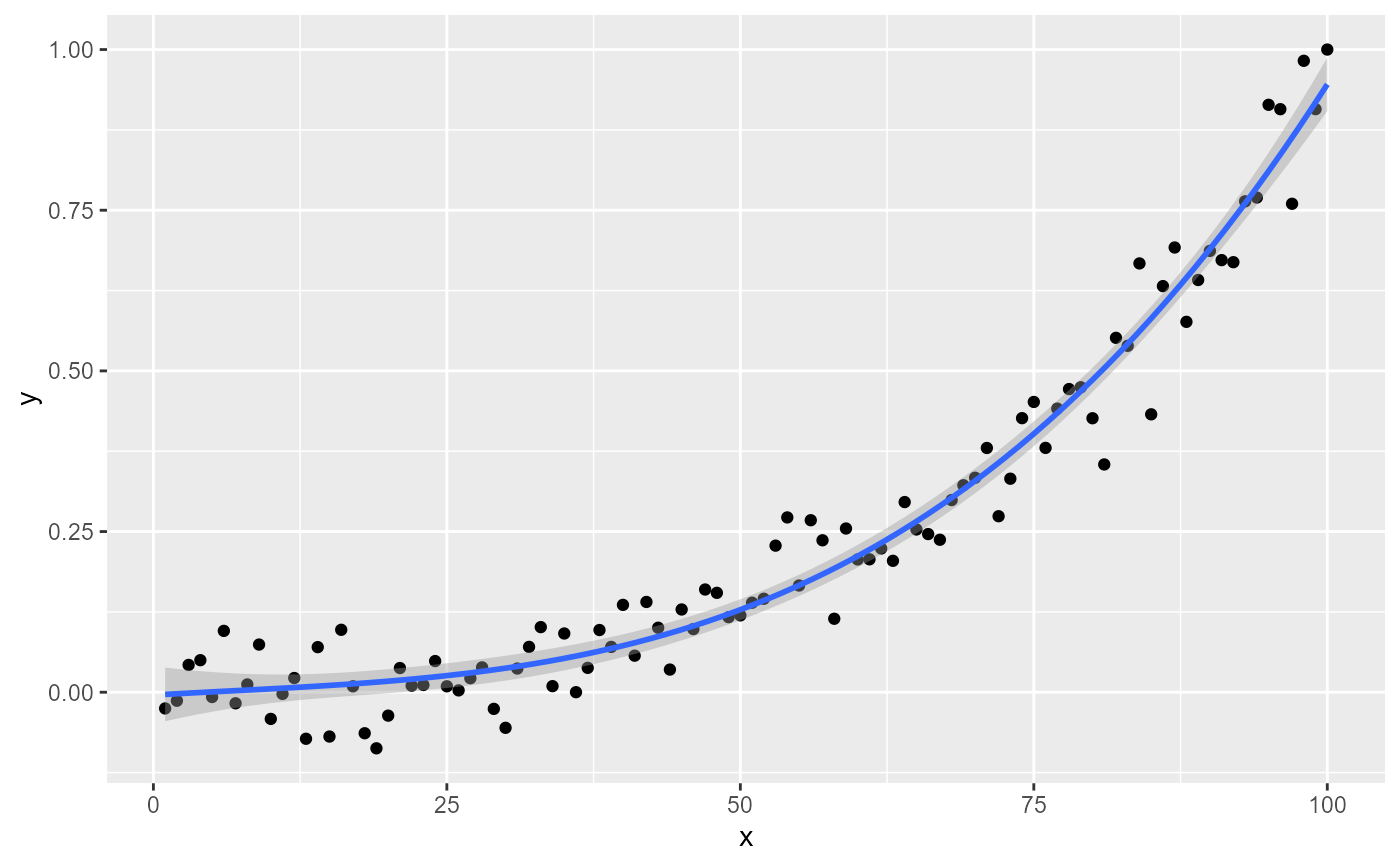
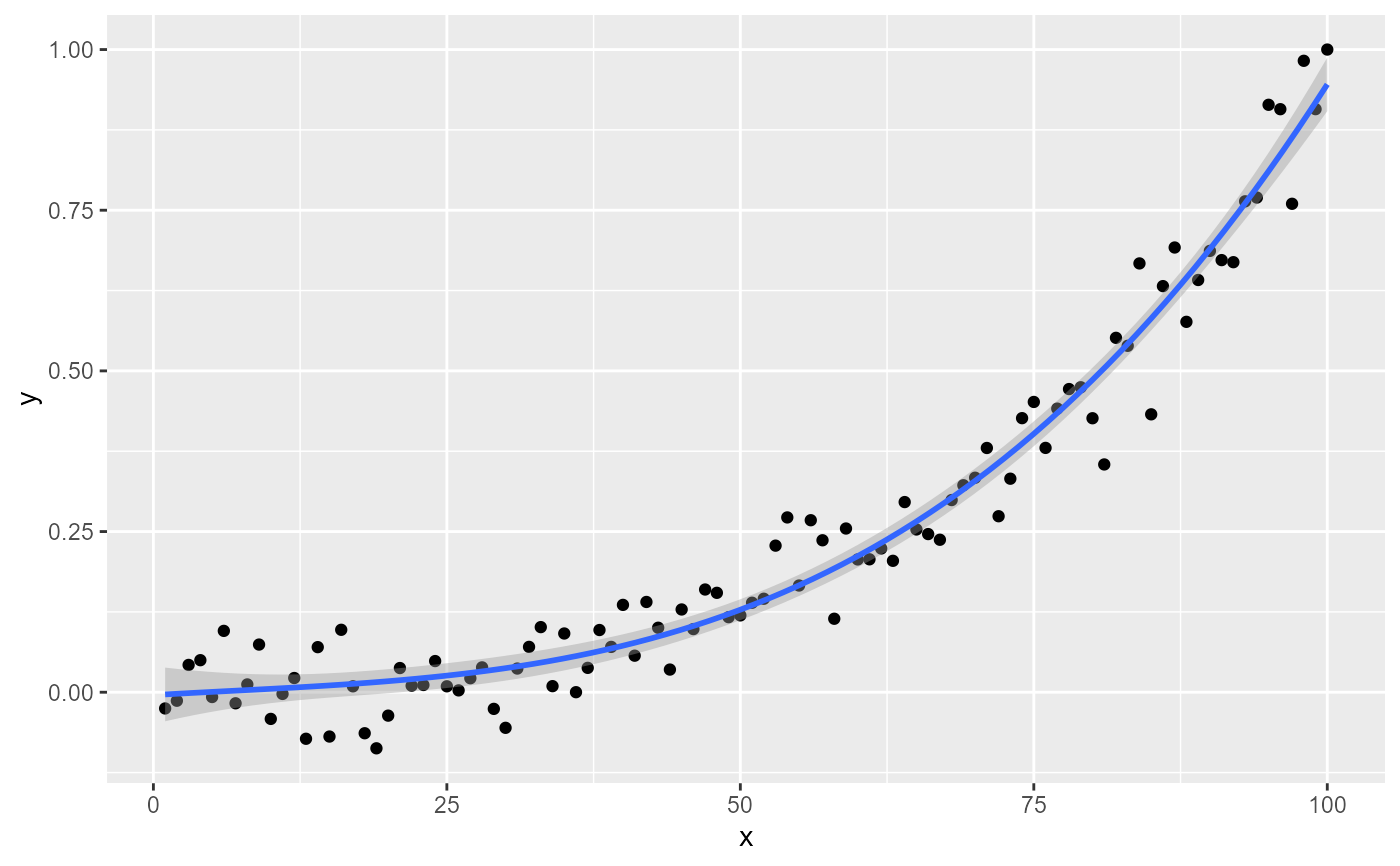
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line() +
stat_poly_eq()
 # no weights
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
# no weights
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
 # other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula)
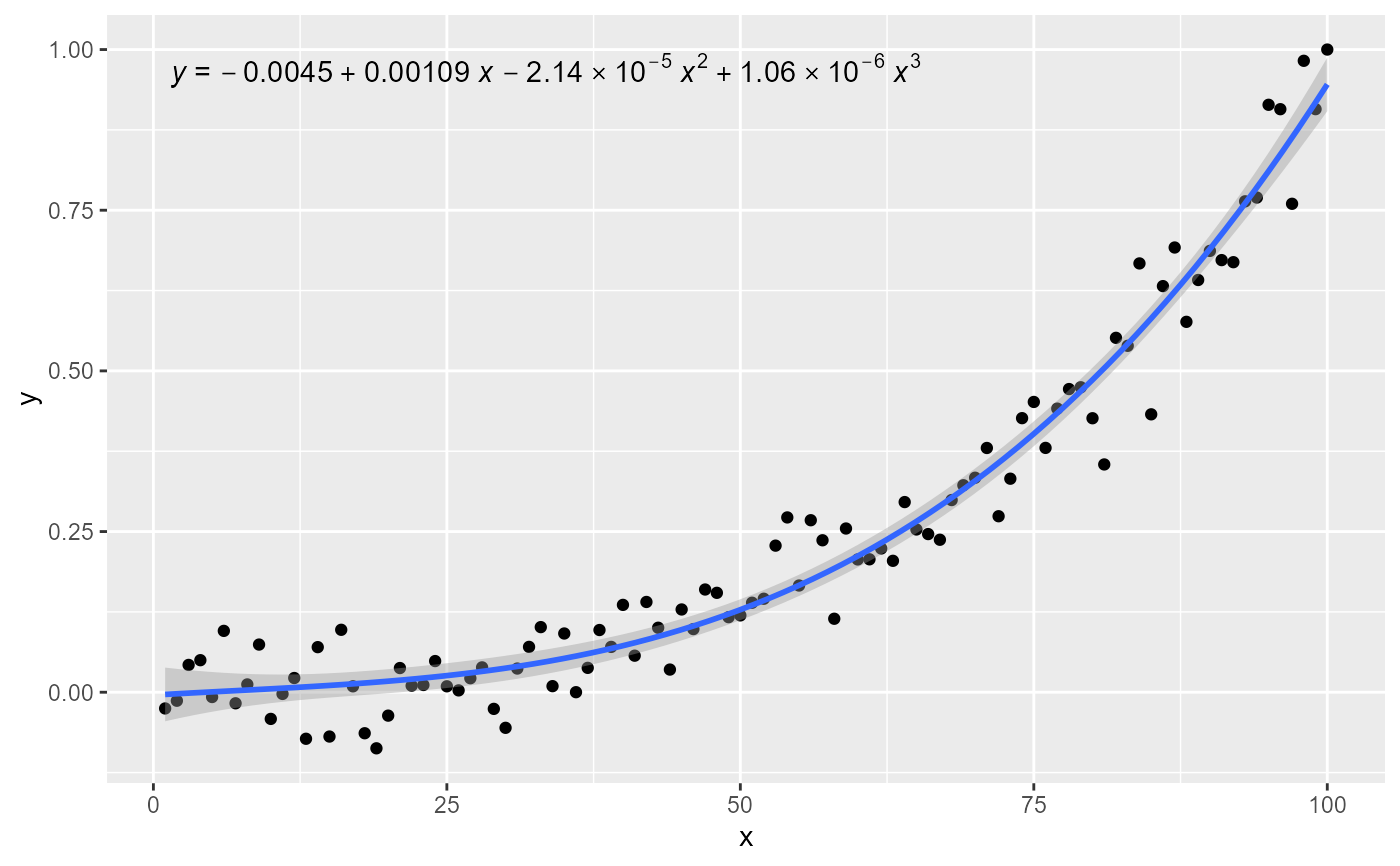
# other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula)
 # other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula, decreasing = TRUE)
# other labels
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq"), formula = formula, decreasing = TRUE)
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq", "R2"), formula = formula)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("eq", "R2"), formula = formula)
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "R2.CI", "P", "method"), formula = formula)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "R2.CI", "P", "method"), formula = formula)
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "F", "P", "n", sep = "*\"; \"*"),
formula = formula)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(use_label("R2", "F", "P", "n", sep = "*\"; \"*"),
formula = formula)
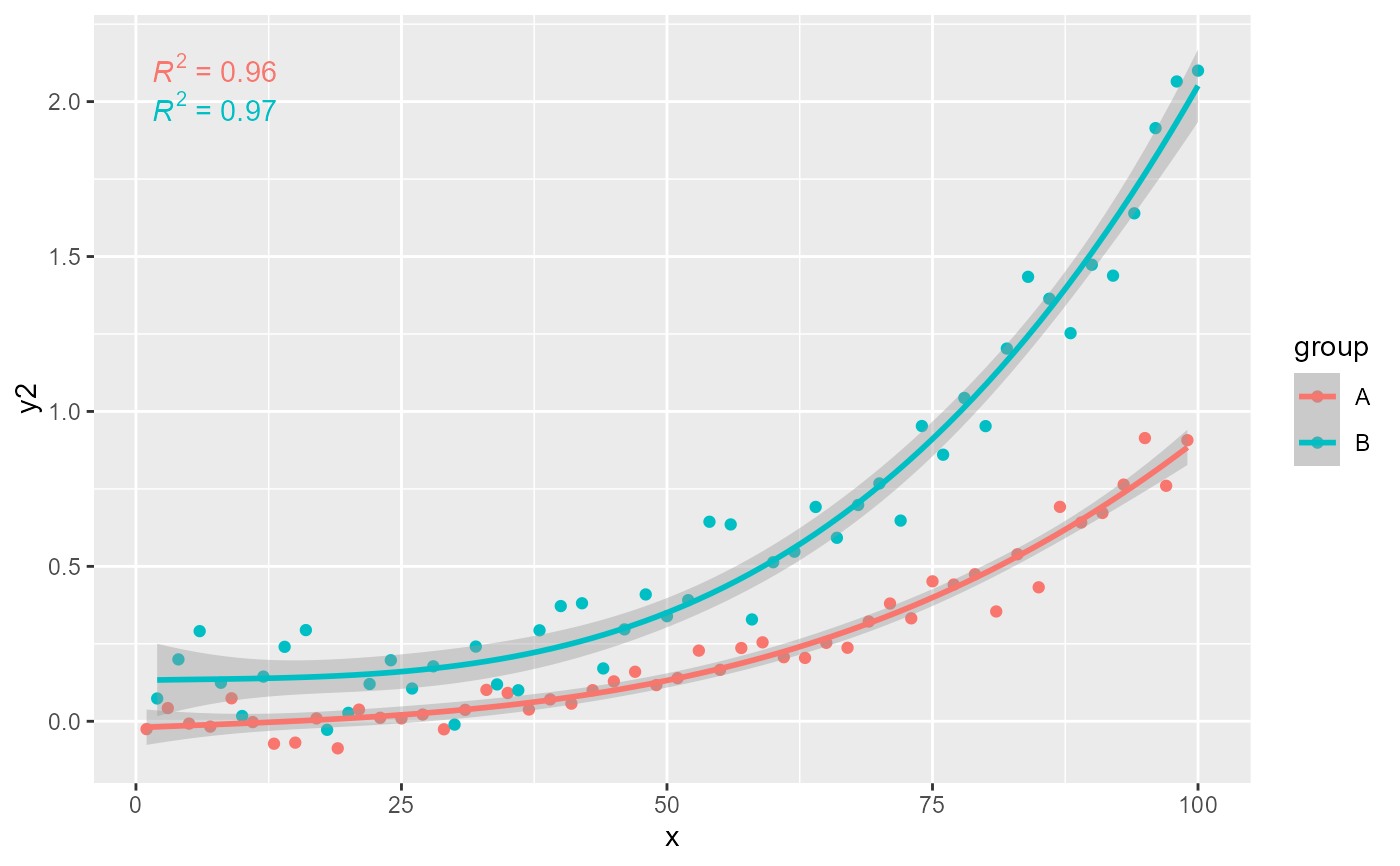 # grouping
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
# grouping
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
 # rotation
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, angle = 90)
# rotation
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, angle = 90)
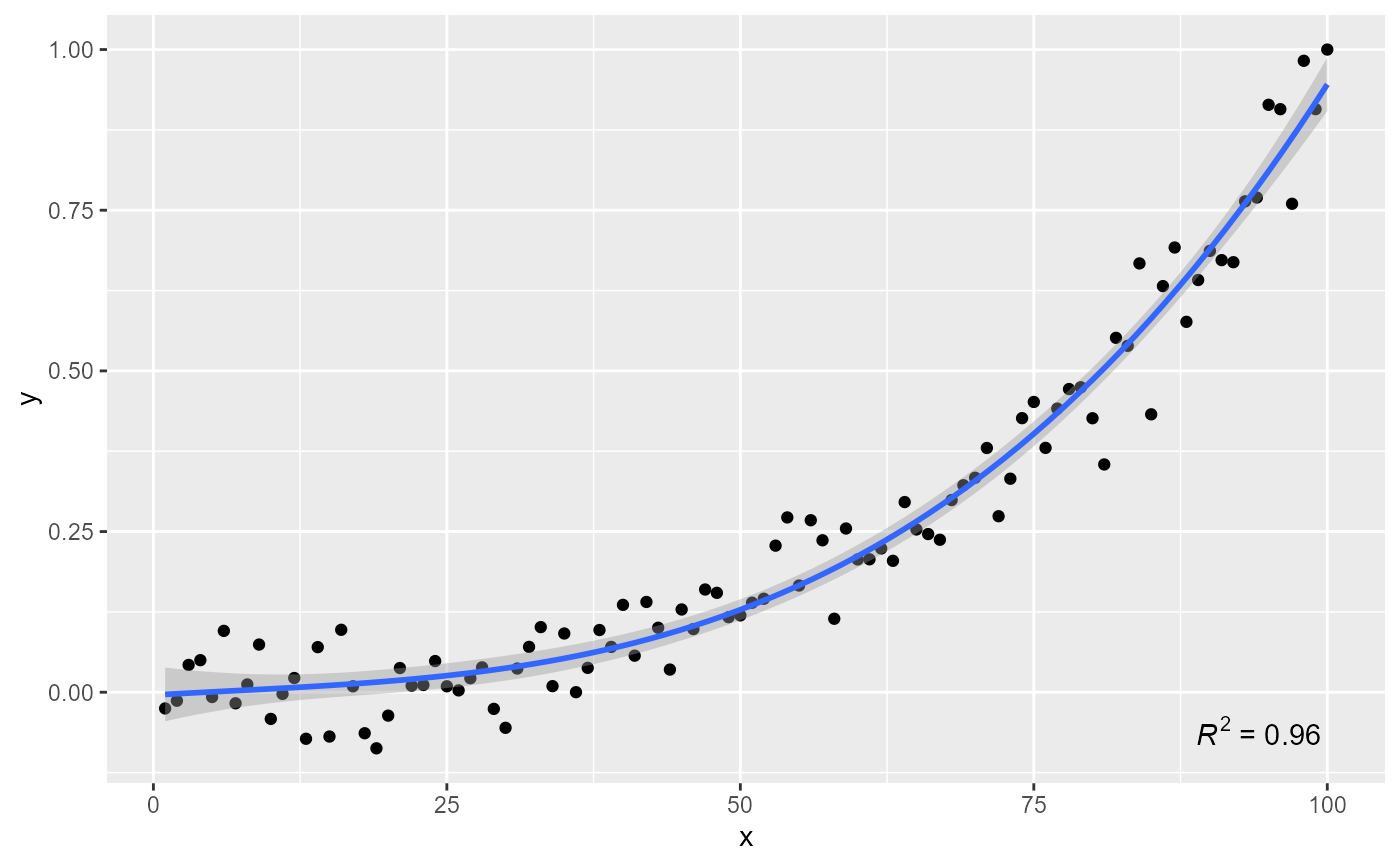 # label location
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = "bottom", label.x = "right")
# label location
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = "bottom", label.x = "right")
 ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = 0.1, label.x = 0.9)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, label.y = 0.1, label.x = 0.9)
 # modifying the explanatory variable within the model formula
# modifying the response variable within aes()
# eq.x.rhs and eq.with.lhs defaults must be overridden!!
formula.trans <- y ~ I(x^2)
ggplot(my.data, aes(x, y + 1)) +
geom_point() +
stat_poly_line(formula = formula.trans) +
stat_poly_eq(use_label("eq"),
formula = formula.trans,
eq.x.rhs = "~x^2",
eq.with.lhs = "y + 1~~`=`~~")
# modifying the explanatory variable within the model formula
# modifying the response variable within aes()
# eq.x.rhs and eq.with.lhs defaults must be overridden!!
formula.trans <- y ~ I(x^2)
ggplot(my.data, aes(x, y + 1)) +
geom_point() +
stat_poly_line(formula = formula.trans) +
stat_poly_eq(use_label("eq"),
formula = formula.trans,
eq.x.rhs = "~x^2",
eq.with.lhs = "y + 1~~`=`~~")
 # using weights
ggplot(my.data, aes(x, y, weight = w)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
# using weights
ggplot(my.data, aes(x, y, weight = w)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula)
 # no weights, 4 digits for R square
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, rr.digits = 4)
# no weights, 4 digits for R square
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula, rr.digits = 4)
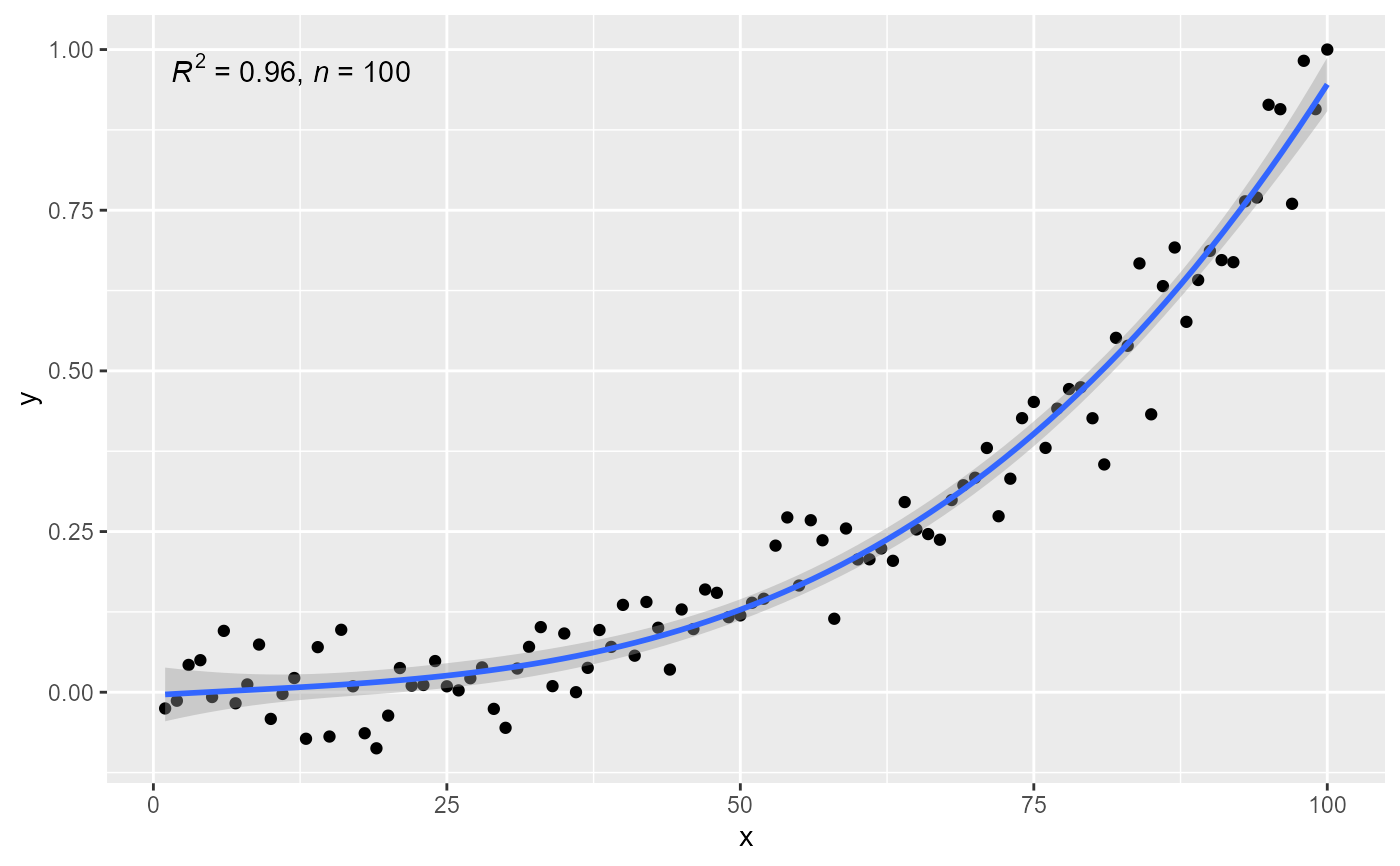 # manually assemble and map a specific label using paste() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = paste(after_stat(rr.label),
after_stat(n.label), sep = "*\", \"*")),
formula = formula)
# manually assemble and map a specific label using paste() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = paste(after_stat(rr.label),
after_stat(n.label), sep = "*\", \"*")),
formula = formula)
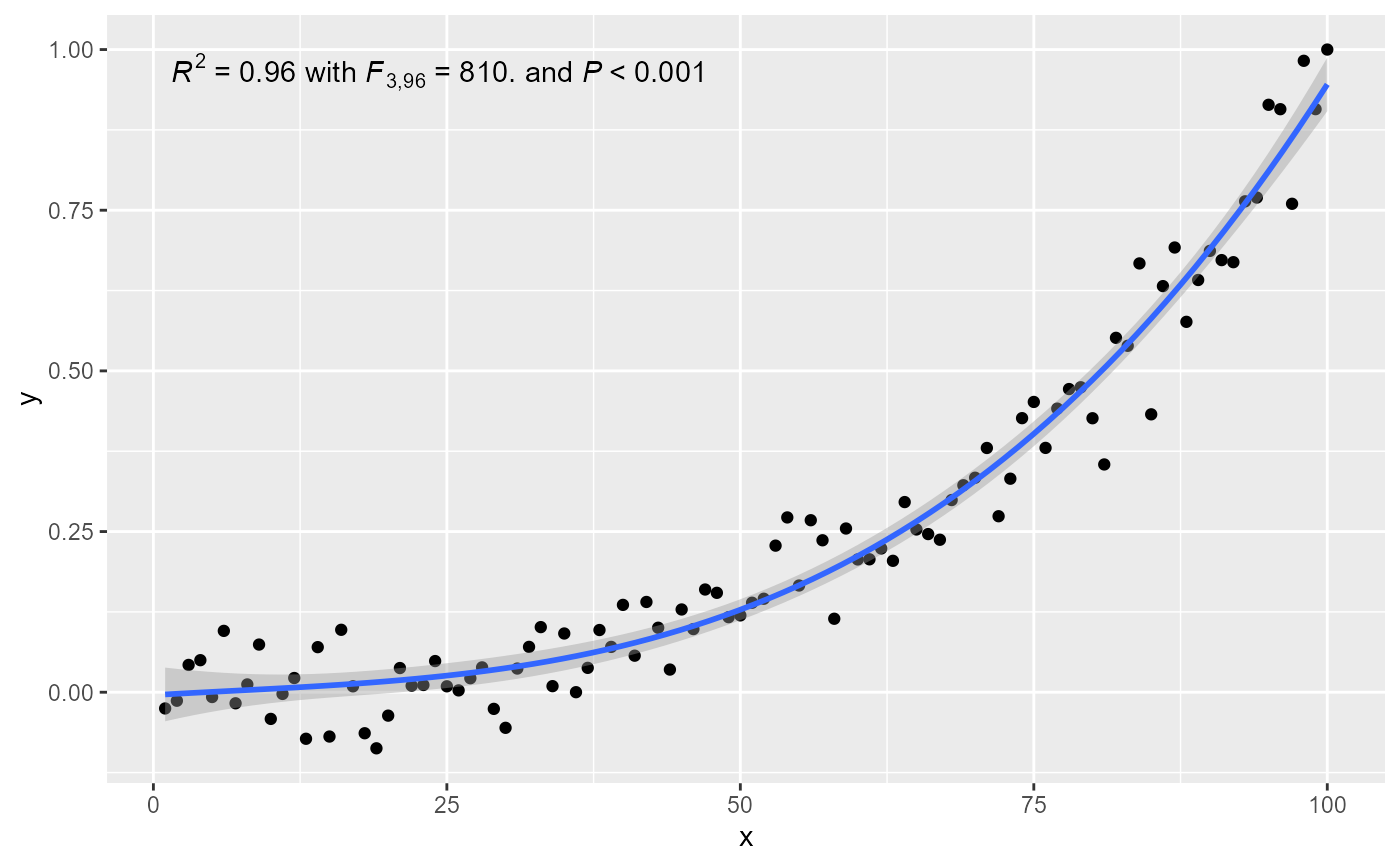 # manually assemble and map a specific label using sprintf() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = sprintf("%s*\" with \"*%s*\" and \"*%s",
after_stat(rr.label),
after_stat(f.value.label),
after_stat(p.value.label))),
formula = formula)
# manually assemble and map a specific label using sprintf() and aes()
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = sprintf("%s*\" with \"*%s*\" and \"*%s",
after_stat(rr.label),
after_stat(f.value.label),
after_stat(p.value.label))),
formula = formula)
 # x on y regression
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula, orientation = "y") +
stat_poly_eq(use_label("eq", "adj.R2"),
formula = x ~ poly(y, 3, raw = TRUE))
# x on y regression
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula, orientation = "y") +
stat_poly_eq(use_label("eq", "adj.R2"),
formula = x ~ poly(y, 3, raw = TRUE))
 # conditional user specified label
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = ifelse(after_stat(adj.r.squared) > 0.96,
paste(after_stat(adj.rr.label),
after_stat(eq.label),
sep = "*\", \"*"),
after_stat(adj.rr.label))),
rr.digits = 3,
formula = formula)
# conditional user specified label
ggplot(my.data, aes(x, y2, color = group)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(aes(label = ifelse(after_stat(adj.r.squared) > 0.96,
paste(after_stat(adj.rr.label),
after_stat(eq.label),
sep = "*\", \"*"),
after_stat(adj.rr.label))),
rr.digits = 3,
formula = formula)
 # geom = "text"
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(geom = "text", label.x = 100, label.y = 0, hjust = 1,
formula = formula)
# geom = "text"
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(geom = "text", label.x = 100, label.y = 0, hjust = 1,
formula = formula)
 # Inspecting the returned data using geom_debug_group()
# This provides a quick way of finding out the names of the variables that
# are available for mapping to aesthetics with after_stat().
gginnards.installed <- requireNamespace("gginnards", quietly = TRUE)
if (gginnards.installed)
library(gginnards)
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group")
# Inspecting the returned data using geom_debug_group()
# This provides a quick way of finding out the names of the variables that
# are available for mapping to aesthetics with after_stat().
gginnards.installed <- requireNamespace("gginnards", quietly = TRUE)
if (gginnards.installed)
library(gginnards)
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group")
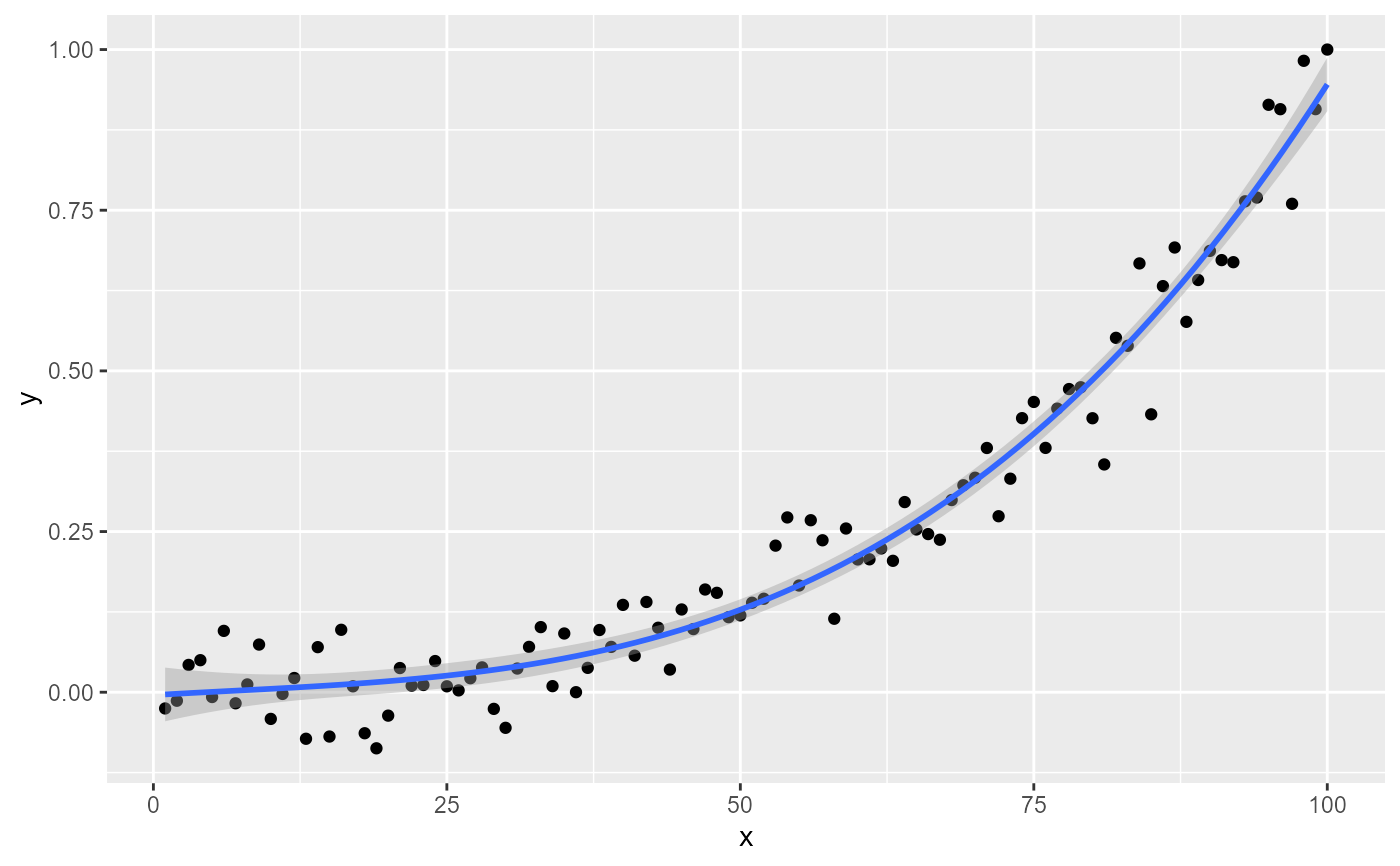 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> knots.label grp.label method.label
#> 1 <NA> -1 "method: lm:qr"
#> coefs
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06
#> coefs.names
#> 1 (Intercept), poly(x, 3, raw = TRUE)1, poly(x, 3, raw = TRUE)2, poly(x, 3, raw = TRUE)3
#> rr.confint.level rr.confint.low rr.confint.high f.value f.df1 f.df2 r.squared
#> 1 0.95 0.9464423 0.9696371 810.484 3 96 0.9620171
#> adj.r.squared p.value AIC BIC n knots knots.se knots.names
#> 1 0.9608301 5.110349e-68 -291.8639 -278.838 100 NA NA NA
#> fm.method fm.class fm.formula fm.formula.chr x
#> 1 lm:qr lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1
#> npcx y npcy PANEL group label orientation
#> 1 NA 1 NA 1 -1 italic(R)^2~`=`~"0.96" x
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
output.type = "numeric")
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> eq.label
#> 1 italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3
#> rr.label adj.rr.label rr.confint.label
#> 1 italic(R)^2~`=`~"0.96" italic(R)[adj]^2~`=`~"0.96" "95% CI [0.95, 0.97]"
#> AIC.label BIC.label
#> 1 plain(AIC)~`=`~"-291.9" plain(BIC)~`=`~"-278.8"
#> f.value.label p.value.label n.label
#> 1 italic(F)[3*","*96]~`=`~"810." italic(P)~`<`~"0.001" italic(n)~`=`~100
#> knots.label grp.label method.label
#> 1 <NA> -1 "method: lm:qr"
#> coefs
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06
#> coefs.names
#> 1 (Intercept), poly(x, 3, raw = TRUE)1, poly(x, 3, raw = TRUE)2, poly(x, 3, raw = TRUE)3
#> rr.confint.level rr.confint.low rr.confint.high f.value f.df1 f.df2 r.squared
#> 1 0.95 0.9464423 0.9696371 810.484 3 96 0.9620171
#> adj.r.squared p.value AIC BIC n knots knots.se knots.names
#> 1 0.9608301 5.110349e-68 -291.8639 -278.838 100 NA NA NA
#> fm.method fm.class fm.formula fm.formula.chr x
#> 1 lm:qr lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1
#> npcx y npcy PANEL group label orientation
#> 1 NA 1 NA 1 -1 italic(R)^2~`=`~"0.96" x
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
output.type = "numeric")
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> rr.label
#> 1
#> coef.ls
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06, 2.268244e-02, 1.935252e-03, 4.440449e-05, 2.890821e-07, -1.981987e-01, 5.630500e-01, -4.819189e-01, 3.650820e+00, 8.433087e-01, 5.747135e-01, 6.309604e-01, 4.255076e-04
#> b_0.constant b_0 b_1 b_2 b_3
#> 1 FALSE -0.004495631 0.001089644 -2.139937e-05 1.055387e-06
#> coefs
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06
#> coefs.names
#> 1 (Intercept), poly(x, 3, raw = TRUE)1, poly(x, 3, raw = TRUE)2, poly(x, 3, raw = TRUE)3
#> rr.confint.level rr.confint.low rr.confint.high f.value f.df1 f.df2 r.squared
#> 1 0.95 0.9464423 0.9696371 810.484 3 96 0.9620171
#> adj.r.squared p.value AIC BIC n knots knots.se knots.names
#> 1 0.9608301 5.110349e-68 -291.8639 -278.838 100 NA NA NA
#> fm.method fm.class fm.formula fm.formula.chr x
#> 1 lm:qr lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1
#> npcx y npcy PANEL group label orientation
#> 1 NA 1 NA 1 -1 x
# names of the variables
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
dbgfun.data = colnames)
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (head):"
#> rr.label
#> 1
#> coef.ls
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06, 2.268244e-02, 1.935252e-03, 4.440449e-05, 2.890821e-07, -1.981987e-01, 5.630500e-01, -4.819189e-01, 3.650820e+00, 8.433087e-01, 5.747135e-01, 6.309604e-01, 4.255076e-04
#> b_0.constant b_0 b_1 b_2 b_3
#> 1 FALSE -0.004495631 0.001089644 -2.139937e-05 1.055387e-06
#> coefs
#> 1 -4.495631e-03, 1.089644e-03, -2.139937e-05, 1.055387e-06
#> coefs.names
#> 1 (Intercept), poly(x, 3, raw = TRUE)1, poly(x, 3, raw = TRUE)2, poly(x, 3, raw = TRUE)3
#> rr.confint.level rr.confint.low rr.confint.high f.value f.df1 f.df2 r.squared
#> 1 0.95 0.9464423 0.9696371 810.484 3 96 0.9620171
#> adj.r.squared p.value AIC BIC n knots knots.se knots.names
#> 1 0.9608301 5.110349e-68 -291.8639 -278.838 100 NA NA NA
#> fm.method fm.class fm.formula fm.formula.chr x
#> 1 lm:qr lm y ~ poly(x, 3, raw = TRUE) y ~ poly(x, 3, raw = TRUE) 1
#> npcx y npcy PANEL group label orientation
#> 1 NA 1 NA 1 -1 x
# names of the variables
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
dbgfun.data = colnames)
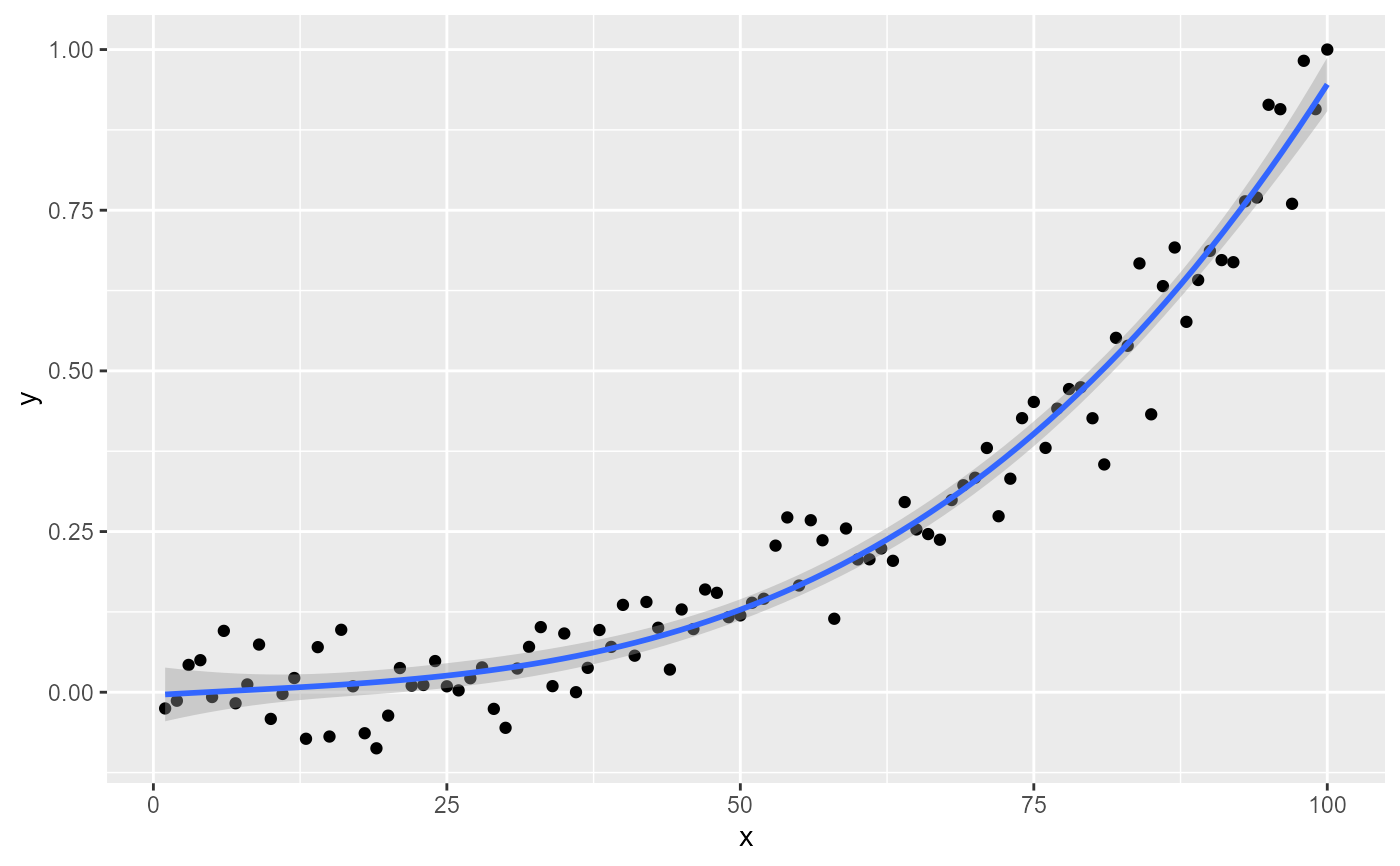 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (anonymous function):"
#> [1] "eq.label" "rr.label" "adj.rr.label" "rr.confint.label"
#> [5] "AIC.label" "BIC.label" "f.value.label" "p.value.label"
#> [9] "n.label" "knots.label" "grp.label" "method.label"
#> [13] "coefs" "coefs.names" "rr.confint.level" "rr.confint.low"
#> [17] "rr.confint.high" "f.value" "f.df1" "f.df2"
#> [21] "r.squared" "adj.r.squared" "p.value" "AIC"
#> [25] "BIC" "n" "knots" "knots.se"
#> [29] "knots.names" "fm.method" "fm.class" "fm.formula"
#> [33] "fm.formula.chr" "x" "npcx" "y"
#> [37] "npcy" "PANEL" "group" "label"
#> [41] "orientation"
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
output.type = "expression",
dbgfun.data = function(x) {x[["eq.label"]]})
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (anonymous function):"
#> [1] "eq.label" "rr.label" "adj.rr.label" "rr.confint.label"
#> [5] "AIC.label" "BIC.label" "f.value.label" "p.value.label"
#> [9] "n.label" "knots.label" "grp.label" "method.label"
#> [13] "coefs" "coefs.names" "rr.confint.level" "rr.confint.low"
#> [17] "rr.confint.high" "f.value" "f.df1" "f.df2"
#> [21] "r.squared" "adj.r.squared" "p.value" "AIC"
#> [25] "BIC" "n" "knots" "knots.se"
#> [29] "knots.names" "fm.method" "fm.class" "fm.formula"
#> [33] "fm.formula.chr" "x" "npcx" "y"
#> [37] "npcy" "PANEL" "group" "label"
#> [41] "orientation"
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
output.type = "expression",
dbgfun.data = function(x) {x[["eq.label"]]})
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (anonymous function):"
#> [1] "italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3"
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
output.type = "text",
dbgfun.data = function(x) {x[["eq.label"]]})
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (anonymous function):"
#> [1] "italic(y)~`=`~-0.00450 + 0.00109*~italic(x) - 2.14%*% 10^{-05}*~italic(x)^2 + 1.06%*% 10^{-06}*~italic(x)^3"
# only data$eq.label
if (gginnards.installed)
ggplot(my.data, aes(x, y)) +
geom_point() +
stat_poly_line(formula = formula) +
stat_poly_eq(formula = formula,
geom = "debug_group",
output.type = "text",
dbgfun.data = function(x) {x[["eq.label"]]})
 #> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (anonymous function):"
#> [1] "y = -0.00450 + 0.00109 x - 2.14 10^-05 x^2 + 1.06 10^-06 x^3"
#> [1] "PANEL 1; group(s) -1; 'draw_function()' input 'data' (anonymous function):"
#> [1] "y = -0.00450 + 0.00109 x - 2.14 10^-05 x^2 + 1.06 10^-06 x^3"